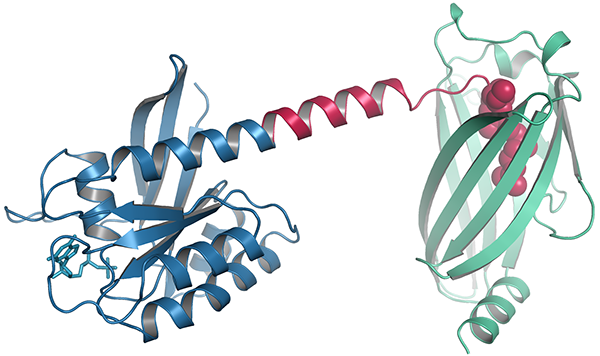
Structure of KRAS4b-PDEδ Complex
, by Dhirendra Simanshu
The three human RAS genes encode four proteins, HRAS, NRAS, and the alternatively spliced forms of KRAS, KRAS4a and KRAS4b. The RAS proteins are identical over their first 86 amino acids and highly homologous for the first 164 (out of 188 or 189 total). Consequently, it is thought that the involvement of different RAS mutants in different cancers is due largely to the C-terminal "hypervariable regions" (HVRs) of the proteins. All the RAS proteins are active only when they are bound to membranes, and they are localized there because their HVRs are modified with lipids, carboxymethyls (one carbon), thioether-linked farnesyl groups (three prenyls, 15 carbons) and thioester-linked palmitoyl groups (16 carbon fatty acid). All the processed RAS proteins have carboxy-terminal cysteines that are both methylated and farnesylated, and HRAS, NRAS, and KRAS4a, but not KRAS4b, have other cysteines in their HVRs that are palmitoylated. Instead, KRAS4b has a polybasic region that is believed to interact with negatively charged head groups of plasma membrane lipids. Thus, KRAS4b is thought to be uniquely dependent on farnesylation and the polybasic region for its membrane localization. Unfortunately, inhibitors of farneslyation proved disappointing as treatments for human cancers1, as KRAS4b was found to undergo alternative prenylation by the addition of a geranylgeranyl (four prenyls, 20 carbons) group instead of a farnesyl group.
The lipid groups on RAS proteins are shielded from the aqueous environment of the cytosol by chaperones as they move to the inner leaflet of the plasma membrane. One of these, PDEδ, has been studied in some detail 2,3. We have recently published the structures of two crystal forms of processed KRAS4b protein complexed with PDEδ4. The structures show for the first time all the amino acids of KRAS4b, including the HVR. Crystal form I reveals structural details of farnesylated–methylated KRAS4b binding to PDEδ, and crystal form II suggests the potential binding mode of geranylgeranylated–methylated KRAS4b to PDEδ. The structures and comparison of the HVRs of KRAS4b from other species show a 5-aa-long sequence motif in KRAS4b that may enable PDEδ to bind both forms of prenylated KRAS4b and thus may have an important and as yet unknown function in normal cells. Surprisingly, absence of the single methyl group that is normally part of the lipid modification of KRAS4b decreased its affinity for PDEδ by 30 fold. The structure revealed the basis for the higher affinity of the carboxymethylated form of the protein. Structure and sequence analysis of various prenylated proteins that have been previously tested for binding to PDEδ provides a rationale for why some prenylated proteins do not bind to PDEδ.
Dhirendra Simanshu is the leader of the RAS Structural Biology Group in the NCI RAS Initiative at the Frederick National Lab.