About Cancer Model Development Centers
The Cancer Model Development Centers (CMDCs) are the NCI-supported contributors to the Human Cancer Models Initiative (HCMI). CMDCs are tasked with producing next-generation cancer models from clinical samples.
The cancer models include tumor types that are rare, originate from patients from underrepresented populations, lack precision therapy, or lack cancer model tools.
Throughout the development process, the CMDCs use stringent internal quality control measures to ensure both clinical and molecular integrity. These models' case-associated clinical, biospecimen, and molecular characterization data are available as a community resource.
The CMDCs are:
- Broad Institute of MIT and Harvard: The co-directors are Jesse S. Boehm, Ph.D. and Keith L. Ligon, M.D., Ph.D.
- Cold Spring Harbor Laboratory: The principal investigator is David A. Tuveson, M.D., Ph.D. This CMDC includes two international subsites: 1) ARC-NET Centre for Applied Research on Cancer, University of Verona, co-led by Aldo Scarpa, M.D., Ph.D. and Vincenzo Corbo, Ph.D. and 2) Hubrecht Institute, led by Hans Clevers, M.D., Ph.D.
- Stanford University: The principal investigators are Calvin J. Kuo, M.D., Ph.D. and Claudia Petritsch, Ph.D.
- Weill Cornell Medical College: The principal investigator is Olivier Elemento, Ph.D.
Locations of Centers and Processing Entities
The map shows locations of the CMDCs (purple): Broad Institute in Cambridge, MA; Cold Spring Harbor Laboratory in Cold Spring Harbor, NY, and their subsites: Hubrecht Institute in Utrecht, Netherlands, and University of Verona in Verona, Italy; Stanford University, Stanford, CA; Weill Cornell Medical College, New York, NY.
Biospecimen Processing Center (green) is the Nationwide Children’s Hospital in Columbus, OH.
Genome Characterization Centers (red) are at Broad Institute in Cambridge, MA, and University of North Carolina in Chapel Hill, NC.
The Genomic Data Commons (pink) is in Chicago, IL.
The Clinical Data Coordinating Center (teal) is at the Information Management Services in Rockville, MD and American Type Culture Collection is in Manassas, VA.
The web developer for HCMI's Searchable Catalog is at the Ontario Institute for Cancer Research (grey) in Toronto, Ontario, Canada.
Quality Controls and Process
The HCMI CMDC-specific model development and characterization pipeline involves multiple institutions. It includes several quality checkpoints, which ensure that the models and case-associated clinical, biospecimen, and molecular characterization data are complete and consistent.
CMDC Development Flowchart
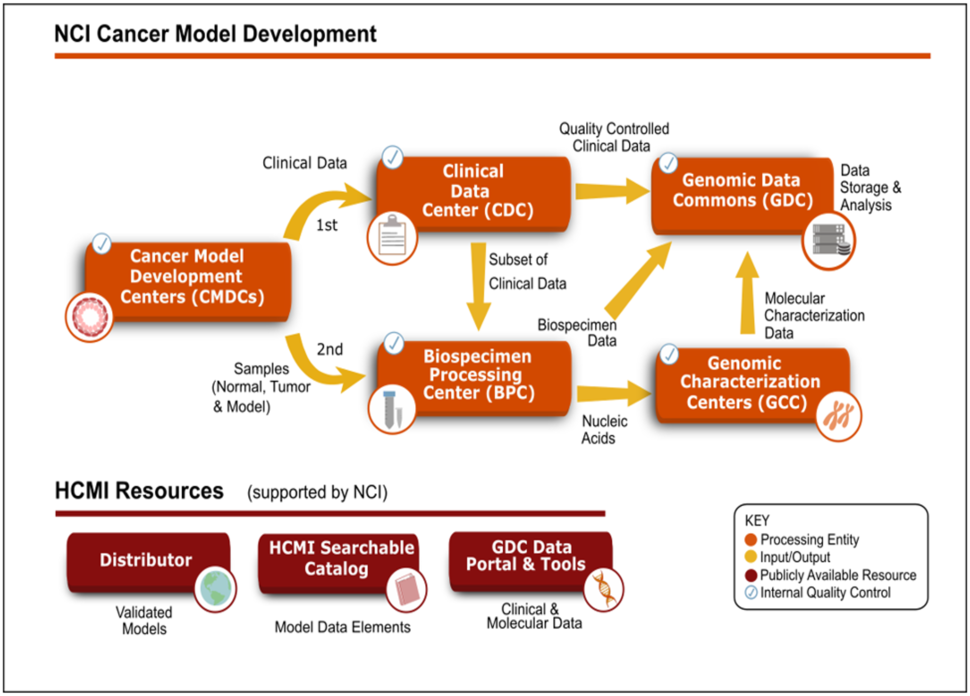
The models are first generated by the CMDCs and clinical data associated with the model's case is submitted to the Clinical Data Center. The Clinical Data Center quality-controlled clinical data for the model's case is sent to the Genomic Data Commons. Samples of normal tissue, parent tumor, and derived model are sent to the Biospecimen Processing Center for isolation of nucleic acids. The quality-controlled nucleic acid samples are sent to the Genome Characterization Centers for whole exome sequencing, whole genome sequencing, and RNA-sequencing. Raw sequencing data is submitted to the Genomic Data Commons. CMDC-validated models and their case-associated genomic and clinical data are provided to the research community when available. More information about each component of the flowchart:
Clinical Data Center (CDC)
The clinical data is submitted to the CDC at Information Management Services Inc. The CDC collects and quality controls the clinical data to confirm patient privacy is protected and to certify that the submitted clinical data conforms to the controlled vocabulary of the case report forms. As part of the clinical data submission process to Genomic Data Commons (GDC), the CDC maps the clinical data to the GDC data dictionary so that users can search and filter by specific clinical data.
Biospecimen Processing Center (BPC)
Tissue samples from patients and derived cancer models are sent to the BPC at Nationwide Children's Hospital. DNA is isolated from normal tissue, parent tumor, and the derived model while RNA is isolated from the parent tumor and derived model using standardized protocols. Internal BPC quality control and genotyping ensure quality and consistency of material for molecular characterization. The BPC submits biospecimen data to GDC and sends isolated nucleic acids to the Genomic Characterization Centers (GCCs) for molecular characterization.
Genomic Characterization Centers (GCC)
The isolated nucleic acids are sent to the GCCs for molecular characterization.
- Broad Institute: 15x whole genome sequencing and 150x whole exome sequencing are performed on DNA from normal tissue, parent tumor (when available), and derived model. Infinium MethylationEPIC DNA Array is performed on DNA from the derived model and a subset of tumors.
- The University of North Carolina: 120 million read RNA sequencing is performed on RNA from the parent tumor and the derived model.
Genomic Data Commons (GDC)
NCI's GDC houses all the clinical, biospecimen, and molecular characterization data. The GDC performs quality control and harmonizes the sequencing data through their analytical pipeline. Harmonized data are available at the GDC Data Portal.
All data is made available using National Institutes of Health policies that protect patient privacy and confidentiality.
Distributor
The models are broadly available through a single third-party distributor, American Type Culture Collection (ATCC). ATCC expands, preserves, quality checks, and globally distributes the models. ATCC provides standard operating procedures, images, and other model-specific culture information to the research community.