HCMI Program Updates and New Features of the Searchable Catalog
, by Eva Tonsing-Carter, Ph.D. and Cindy Kyi, Ph.D.
The Human Cancer Models Initiative (HCMI) is an international collaboration between the National Cancer Institute (NCI), Cancer Research UK (CRUK), Wellcome Sanger Institute (WSI), and foundation Hubrecht Organoid Technology (HUB). The goal of the HCMI is to create up to 1,000 next-generation cancer models from patient tumors that are annotated with clinical, biospecimen, and molecularly characterization data.
The HCMI models and their case-associated data are available to researchers as a community resource. Existing models may be queried at the HCMI Searchable Catalog, a continuously updated resource of available HCMI models. Since our last e-News article in August 2020, new models have been generated; bringing the total number of currently available models to 148 models from 18 different primary sites including brain, skin, colon, pancreas, lung, extrahepatic bile duct, and more. Of the 148 models, there are currently 10 models from five cases, which have multiple models derived from unique anatomic sites from the same patient. These include models derived from primary and metastatic or multiple metastatic tumors.
Quality-controlled and harmonized clinical, biospecimen, and molecular characterization data for 63 models from 62 cases are accessible as of February 16, 2021 at NCI’s Genomic Data Commons (GDC) Data Portal. Excitingly, one of these cases has multiple models derived from primary and metastatic ampulla of Vater cancer. The molecular characterization data includes RNA-sequencing, whole genome sequencing, whole exome sequencing, copy number variation, and masked somatic MAF data. Epigenetic data from a subset of models and their originating tumors will be available in the future. Data from new cases are being released as they become available. To access the data, visit the GDC’s Data Portal. dbGaP approval is required for accessing controlled-access data. The clinical, biospecimen, and masked somatic MAF data are open-access and do not require approval to access. Visit the Accessing HCMI data webpage for more detailed information on how to access the HCMI data.
The HCMI Searchable Catalog has been updated with new interactive features for enhanced user experience in sorting and browsing the available HCMI models. New cancer types, new search features, multiple models from some cases, and open-access masked somatic MAF data for a subset of models have been added to the Catalog.
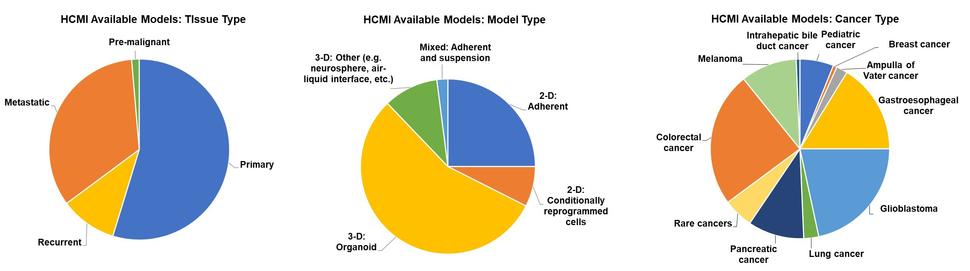
A summary of currently available models by cancer type, model type, and tissue type is described below (Figure 1).
Landing page
The Searchable Catalog landing page contains a main viewing table with a list of all available models within the Catalog (Figure 2). The Catalog has been streamlined with user interface enhancements including updated color scheme, increased text contrast, and helpful links in the footer. Available open-access masked somatic MAF data generated at the GDC have been integrated into the Catalog as a searchable element. Users can also search by gene, specific somatic variants, and type of research somatic variant (e.g. missense, nonsense, etc.). As the model data elements are selected, the results listed within the main viewing table (shown within the red border in Figure 2) will update dynamically. There are a few HCMI cases where multiple models were derived from unique sites from the same patient, and “Has Multiple Models” has been added as a search option to identify these cases.
The top panel on the landing page shows interactive circular graphs which users may use to filter the models by primary site, availability of multiple models, 2D versus 3D growth, and most frequently mutated genes by clicking on various colors on the graphs. Hovering over different colors within a graph will reveal relevant information.
Within the main viewing table, columns indicating the numbers of mutated genes, research somatic variants from the open-access masked somatic MAF data, clinical sequencing variants, and histopathological biomarkers have been added. Users can customize the columns shown by selecting data of interest from the “COLUMNS” dropdown menu. Users can sort the data displayed within the table in ascending or descending order by clicking on the headers of their choice within the main viewing table. The selected data can be exported using the “EXPORT ALL” function, saved as a .tsv file, and opened in Excel or similar program.
Users may conduct multiple searches and save models of interest by selecting the checkbox next to the model name. The selected models are added to “My Models List” and their associated data can be downloaded as a .tsv file for later usage.
Navigating individual model pages
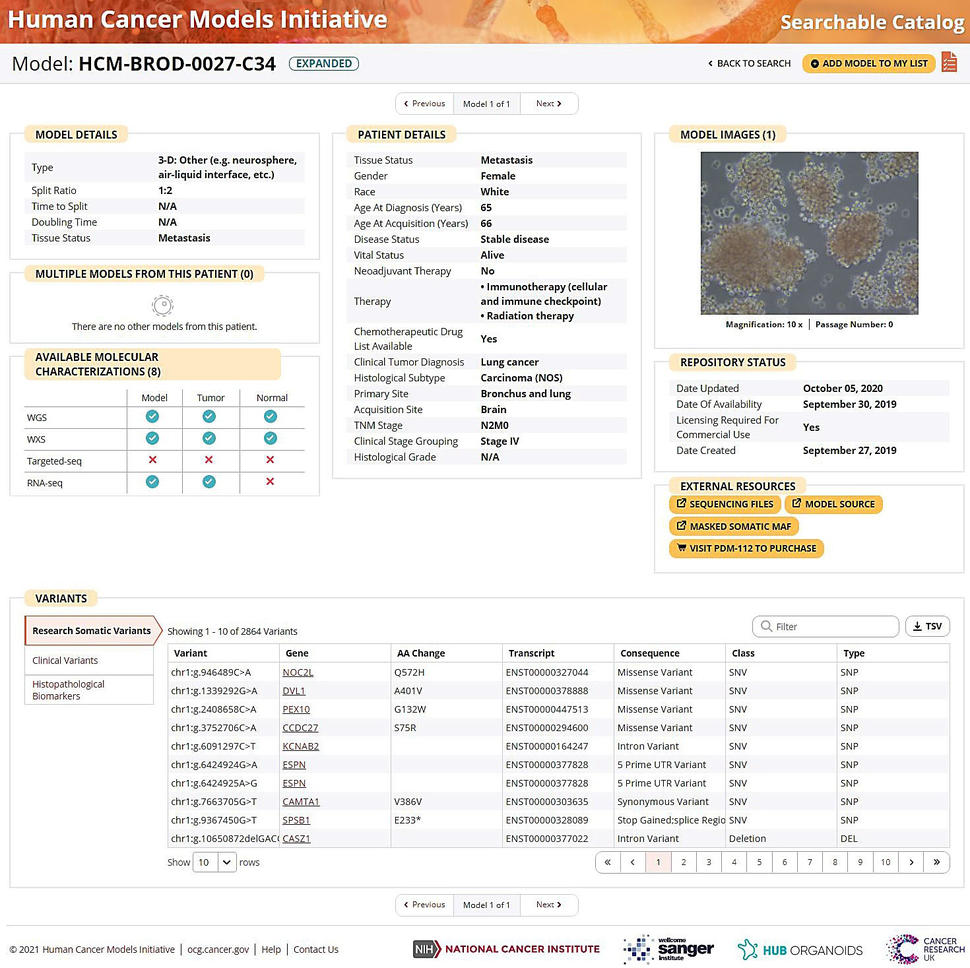
To view details of individual models, users may click on a model name of interest, and they will be routed to the model page (Figure 3).
The user interface of the individual model pages have been rearranged to allow users to quickly identify all the available information for any model. Data on individual model pages include model details such as the type, split ratio, tissue status, etc. and links to multiple models generated from the same patient (if available). The molecular characterization data types that are accessible at the GDC, patient details, model images in culture, and links to external resources such as the model distributor and the GDC are also displayed.
At the bottom of the page, the clinical sequencing and histopathological biomarkers provided from the medical records are listed (when available). The available open-access masked somatic MAF data generated at the GDC are integrated into the HCMI Searchable Catalog. Users can search, sort, and download the variant data by clicking the download .tsv icon. The open-access masked somatic MAFs are highly processed to remove lower quality and potential germline variants. If omission of true-positive somatic mutations is a concern, we recommend accessing controlled-access MAFs, which requires user certification through dbGaP; visit Accessing HCMI Data for more information.
Users are encouraged to visit the Catalog frequently as new models and their associated data are being added to the HCMI Searchable Catalog as they become available. Users who have questions about HCMI, the models or the data are encouraged to visit the HCMI FAQ page or visit the HCMI program. A user guide is available to help users navigate the Catalog.
The HCMI models together with their case-associated data provide a robust resource for the research community. The models can be used in a variety of research endeavors to support precision oncology from investigating pathways that influence tumor initiation and progression to studying drug resistance and drug response.