CTD² PANcancer Analysis of Chemical Entity Activity (PANACEA) DREAM Challenge
, by Eugene F. Douglass Jr., Ph.D., Bence Szalai, Ph.D., Robert J. Allaway, Ph.D., and Andrea Califano, Ph.D.
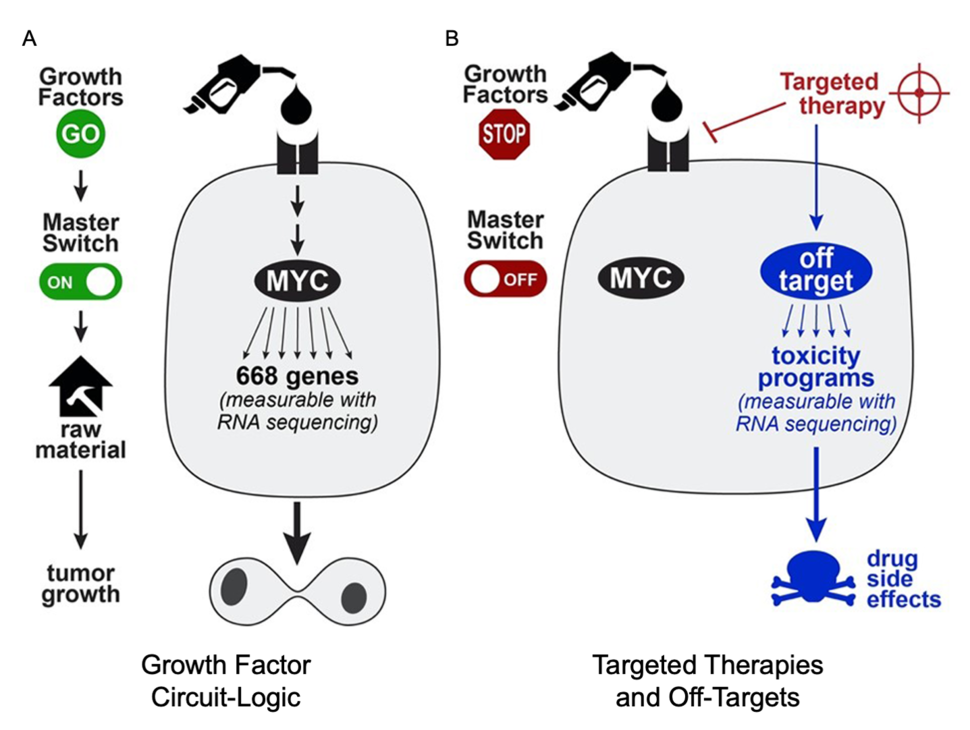
During tissue-repair or wound-healing, growth factors are released to signal the cells to turn on “master switches” like the transcription factor MYC for cellular proliferation. These “master switches” (or “master regulators”) turn on hundreds of genes which serve as the raw materials to build copies of the cell. In cancer, DNA mutations often cause these growth circuits to become stuck on the “on” position, causing uncontrolled growth of the cancer cell (Figure 1A).
Targeted therapies are types of drugs that block specific components of growth circuits, forcing them to return to an off state that halts cellular proliferation and tumor growth. Unfortunately, targeted therapies can sometimes have other effects on the cell (off-targets) which can turn on other programs associated with drug side-effects and toxicity (Figure 1B). Identifying the off-targets of drugs is experimentally challenging, as it requires measuring the drug-target interactions on the proteome scale. As off-targets can drive adverse effects as well as therapeutic effects, identification of the whole target spectrum of targeted therapies is very important.
To better understand the off-target programs of cancer therapies, we built an RNA-sequencing (RNA-Seq) database called PANACEA (PANcancer Analysis of Chemical Entity Activity), which measures the mRNA expression level of all 20,000 protein-coding genes that are turned on or off by 400 FDA-approved or late stage clinical cancer drugs. These RNA-Seq datasets can provide hints about the changes in activities of key signaling pathways of the cells, including those associated with tumor growth (Figure 1A) and off-target programs (Figure 1B).
As PANACEA includes data from several cancer cell lines, it makes possible to observe the tissue of origin-based differences in these transcription programs. Beside RNA-Seq, the cell-death inducing abilities of the investigated drugs were also measured, at different concentrations, to identify the sensitive and resistant cell lines for each drug. This large-scale dataset gives excellent possibility to understand the key mechanisms of drug sensitivity, resistance, and off-target toxicity. Because of the large number of measurements (400 drugs x 21 cell lines x 20,000 genes), this database is suited to exploration using data science and machine learning tools (manuscript in preparation).
With this in mind, we hosted a community challenge with the DREAM Challenges initiative to identify the best machine learning methods to identify off-targets from 30 of the most common clinically-used targeted therapies. We provided the RNA-Seq data and drug sensitivity data from 11 cell lines for the 30 test molecules, from which the participants had to predict the targets (both on- and off-targets) for each molecule. The participants used public databases of RNA-Seq, drug sensitivity measurements, and drug targets to build their computational models. The name and chemical structure of the 30 test molecules were unknown to the participants to ensure that the developed methods were unbiased.
Over two months, 21 teams contributed 86 models, of which 39 (45%) showed statistically significant enrichment of true off-targets within druggable proteins. The best performing methods leveraged a variety of approaches including fundamental chemical and transcriptomic similarity approaches and more sophisticated deep-learning methodologies. This study lays the foundation for future integrative analyses of pharmacogenomic data, reconciliation of polypharmacological effects in different tumor contexts, and insights into network-based assessment of context-specific drug mechanism of action. More information about the results of the challenge can be found at the Challenge landing page and in our recent preprint on bioRxiv.1
- Douglass Jr. E, Allaway RJ, Szalai B, et al. A Community Challenge for Pancancer Drug Mechanism of Action Inference from Perturbational Profile Data. bioRxiv. 2020 December 23. doi:10.1101/2020.12.21.423514