NCI Genomic Data Commons: A Community Resource for Cancer Research
, by Louis M. Staudt, M.D., Ph.D.
Data sharing plays a pivotal role in precision oncology. That was the impetus for building the Genomic Data Commons (GDC), an information system that we at the NCI started conceptualizing back in 2014. Two years later, together with our partners at the University of Chicago and the Ontario Institute for Cancer Research, we laid out our vision for the GDC: a community resource that would offer uniformly processed genomic data and facilitate all types of researchers in their precision oncology work.
In a pair of publications this week, the GDC team describes what we’ve built and accomplished so far, including the interactive data system and the genomic data processing pipelines that we’ve assembled and continue to refine.
The GDC: A Killer App, Not a Zip Drive
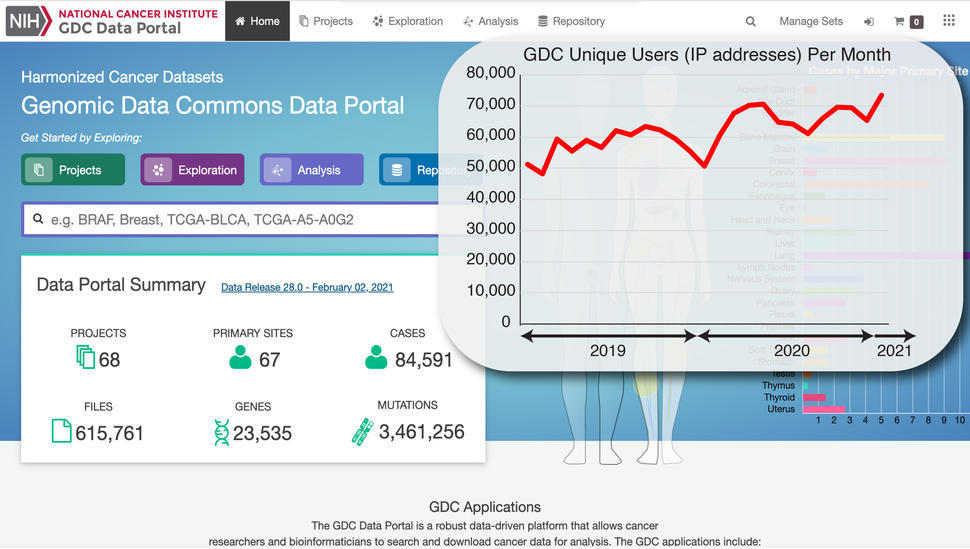
The GDC currently offers over 3.3 PB of data from over 65 projects. The GDC is unique in the world of cancer research in offering a large array of primary and derived genomic data types, including whole genome sequencing, whole exome sequencing, transcriptome sequencing (RNA-seq), microRNA sequencing, DNA methylation analysis, DNA copy number analysis, ATAC-Seq to identify genomic regions with open chromatin, and single nucleus/single cell RNA-seq. In addition, the GDC also includes a rich set of clinical data associated with each case.
But the GDC is more than just a data repository. Each data set that is submitted to the GDC is subject to rigorous genomic and clinical data quality standards and is processed according to a common set of bioinformatic pipelines before being made available to the research community. This allows for a meta-analysis of data in the GDC, in which the user can construct custom cohorts of cases drawn from more than one study. Thus, the GDC is a research platform as well as a secure and stable home for valuable cancer genomic data.
The GDC is continuously adapting its analytical methods to conform with recent advances. Often there is no “right answer” when it comes to analyzing genomic data, and in these cases the GDC reports the results of several analytical pipelines.
For example, the GDC currently predicts somatic mutations using five different algorithms and provides all of these to the user. For many cancer mutations, all pipelines agree, but difficult edge cases may be dealt with better by one algorithm that by the others. Particularly vexing are mutations in tumor subclones that have a low allelic fraction and mutations that involve complicated insertions and deletions of base pairs. In short, as the bioinformatics of cancer evolves, so does the GDC.
Spawning a Data Ecosystem of Applications
GDC data can be consumed in a variety of ways depending on user preference and expertise. Data are available for direct download from a web browser, a command-line interface, through an application programming interface (API), or through several NCI Cloud Resources that operate in commercial cloud workspaces.
The GDC offers a suite of web-based tools to help researchers analyze GDC data even if they do not have an extensive background in bioinformatics or access to computational resources. These Data Analysis, Visualization, and Exploration tools (called DAVE for short) allow users to build custom cohorts of cancer samples, browse mutations, copy number and gene expression levels for individual genes, and create survival curves using the associated clinical data, all in real time without downloading any data.
“We hope by creating the GDC platform with an API that provides access to all its underlying data, we will encourage others to develop systems and applications to analyze GDC data and foster cancer research—effectively making the GDC the hub of an ecosystem of applications supporting cancer genomics research,” says Robert L. Grossman, Ph.D., the GDC’s Principal Investigator.
Harmonization for All (Types of Data)
12 years of The Cancer Genome Atlas (TCGA) taught us not only about cancer biology, but also how to generate and process large quantities of cancer genomic data, even as the molecular analysis platforms constantly evolve. TCGA researchers discovered pretty early on that major batch effects occur if data are aligned to different reference genomes, mapped to different gene models, or processed using different analytical algorithms.
The GDC’s answer to this has been to “harmonize” the data—to process the data in a uniform manner, allowing researchers to more readily compare data acquired from different studies.
Over the past 7 years, the team has worked to develop pipelines for DNA-Seq (whole genome, whole exome, and targeted sequencing), RNA-Seq (mRNA and miRNA), and copy number and DNA methylation array data. A lot of work goes into constructing the pipeline to support any single experimental type—and the GDC supports over 40 pipelines (but who’s counting). The major workflows described in the papers, including GDC developed software tools, Docker files, and pipelines in the Common Workflow Language, can be found in GitHub.
“One of the challenges is working with pipelines that were often developed to be run manually by a single lab. These pipelines often need to be refactored and integrated into an automation framework so they can be run at the scale required by the GDC. We make all of the repackaged pipelines public, so that the broader research community may reap the benefits of all these efforts,” says Dr. Grossman.
Genomic Data Sharing is Trending
In our 2016 perspective describing our vision of the GDC, we wrote “the principles and practice of precision oncology will be accelerated by sharing data from thousands of patients with cancer.” Today, we have data from over 80,000 patients in the GDC.
Of course, the GDC will continue to add more cases and new types of genomic data. We are enriching the GDC with genomic datasets that have comprehensive clinical annotations, including treatments and outcome, environmental exposures and other epidemiologic data, other co-morbid conditions, clinical laboratory tests and more.
The GDC already has longitudinal genomic and clinical data from the Multiple Myeloma Research Foundation’s CoMMpass clinical trial and the Leukemia and Lymphoma Society sponsored Beat AML clinical trial. Data from NCI’s Exceptional Responders initiative, the NCI-MATCH precision medicine trial, and other completed NCI clinical trials will appear in the GDC shortly, along with follow up data for the American Association for Cancer Research's Project GENIE. Another notable program is Count Me In, which has contributed data from patients with rare and metastatic cancers.
Many of the GDC’s data sets have linked radiologic and histologic images, which will enable machine learning technologies to improve the precision and reproducibility of cancer diagnosis. Links of GDC genomic data to proteomic datasets, such as in NCI’s CPTAC program, will enable researchers to discover associations between genomic abnormalities and functional regulatory pathways, and to devise genomically inspired therapeutic trials.
Though we’ve been building and adapting the GDC for the last seven plus years, the endeavor still feels fresh and vital to progress against cancer. Usage of the GDC by cancer researchers around the world has been vigorous and sustained, justifying our original hope and expectation that “if we build it, they will come.”