Charting Out New Mysteries with a Tumor Atlas for Small Cell Lung Cancer: Researcher Interview with Charles Rudin and Dana Pe’er
, by Peggy I. Wang
Cancer is a complex entity involving the interplay of malignant cells, cells in the local tissue microenvironment, and immune cells. The disease also changes over time, progressing through a series of transitions, from pre-cancer to malignancy to metastasis. To map out these intracacies in rich detail, NCI established The Human Tumor Atlas Network (HTAN) as part of the Cancer Moonshot initiative.
While The Cancer Genome Atlas (TCGA)—NCI’s flagship cancer characterization program—produced rich “snapshots” of 33 cancer types, many cancer types have still not been thoroughly characterized. Importantly, many cancers still carry poor prognosis and seem to require the detailed time- and space-specific maps proposed by HTAN for clinical progress to be made.
One recent study from HTAN takes a deeper look at small cell lung cancer, a poorly understood and deadly form of lung cancer. I spoke to the two lead researchers of the study from Memorial Sloan Kettering Cancer Center (MSKCC): Dr. Charles Rudin, physician-scientist, and Dr. Dana Pe’er, Chair of MSKCC’s Computational and Systems Biology Program. The single-cell sequencing and clinical data from their study is available at the HTAN Data Portal.
Peggy I. Wang: To set the context, could you describe the Human Tumor Atlas Network? How is it different from TCGA?
Dana Pe’er: HTAN and TCGA are ambitious in different ways. While TCGA created an enormous compendium of “bulk” genome sequencing data across cancers, the goal of HTAN is to describe cancer processes at the much higher resolution of single cells and to incorporate the dimensions of space and time.
To make this possible, the HTAN pursues diverse new technologies and computational methods—some of which are being developed as the project progresses—with the aim of integrating several data types into biologically interpretable atlases.
Rather than taking a census of the major cancers, HTAN has so far focused on a set of vignettes describing cancer transitions, such as pre-cancer to cancer, therapeutic vulnerability to resistance, and local to dispersed cancer, which is what our center studies. But similar to TCGA, creating a large-scale data resource for the community and sharing new methods is central to HTAN.
PIW: What were some of the major technical hurdles you had to overcome for building a single-cell atlas?
DP: Work in the first two years of HTAN included a lot of assay optimization for clinical tissue–how to get high-quality data with good cell-type representation from fine needle aspirates, autopsy tissue and other challenging sources. Many of the multiplexed spatial technologies are still immature and require tweaking and benchmarking. There is still huge potential for new methods to improve analysis of single-cell and multiplexed imaging data.
Charles M. Rudin: For this project, obtaining the samples in an appropriate way for characterization was a big hurdle! Small cell lung cancer (SCLC) is typically not surgically resected, partly because it tends to spread early, so the typical approach is systemic chemotherapy treatment with radiation. So historically, we haven't had substantial tumor resections of SCLC to study in the laboratory.
We collaborated with our interventional radiologists and others who do biopsies of SCLC to get tissue that we can process very quickly for single-cell characterization. The samples need to be processed within hours, so we had people who were dedicated to getting biopsies from the bronchoscopy suite to the lab and processed. It was a team effort to get this done!
PIW: You mentioned developing methods as the project progresses, which sounds similar to how TCGA proceeded. Can you elaborate some more on the development of single-cell methods?
DP: The analysis of single-cell data is not turnkey; it demands that computational biologists work closely with biological domain experts and get a good “feel” for the right parameter and filtering decisions. The decisions that maximize the extraction of real information are different for every data set.
We are still far from getting ideal quantitative descriptions for important biological concepts such as cell-cell similarity and plasticity; we need better tools for capturing rare cell states, delineating active gene programs that drive biological processes, and integrating data across cohorts and across modalities.
On the imaging side, there are major limits on our ability to carry out basic data processing steps such as cell segmentation, assigning molecular features to cells, noise thresholding, and much more! We need better tools to integrate imaging and sequencing data and to take advantage of what spatial information can tell us.
PIW: TCGA characterized around hundreds of samples per tumor type. How many samples are necessary to build an informative and reliable single-cell picture of a cancer, especially one that has a heterogeneous makeup and might vary greatly and between patients?
CMR: I think the difference is the depth at which you analyze any given sample. With single-cell analysis, you are getting tens of thousands of data points per tumor. We still need to do more tumors to understand the landscape and the heterogeneity of the cellular composition of SCLC; twenty-some cases isn't enough! But we deeply interrogate those cases and we get a tremendously detailed view of the composition of those cases.
Actually, with single-cell profiling you get more data from the tumors than with the bulk sequencing of TCGA, and you also get a very different type of data. It’s complementary data because you're looking at a different aspect of the biology.
PIW: Would you be able to take the bulk expression data like what was gathered from TCGA and deconvolute the different cell types and states?
DP: Some powerful computational tools now exist for deconvolving bulk data into constituent cell types, but their performance can’t compare with having high-quality single-cell data.
This is especially true for discovering cell populations, cell states and fate trajectories that we didn’t know anything about. More often, what we aim to do is generate mechanistic hypotheses from HTAN data, and then query TCGA cohorts in a way that leverages their size and diversity.
PIW: Can you explain what do you mean by cell “state”?
CMR: A cell state is a description of what a cell is actually doing, what kind of role that cell is playing. Some cells may express genes that suggest interactions with particular other cells in the tumor microenvironment, or suggesting they might be progenitors of other differentiated cells within the tumor.
PIW: What techniques in addition to single-cell RNA sequencing (scRNA-seq) did you use to help build the atlas? How does it complement the scRNA-seq data?
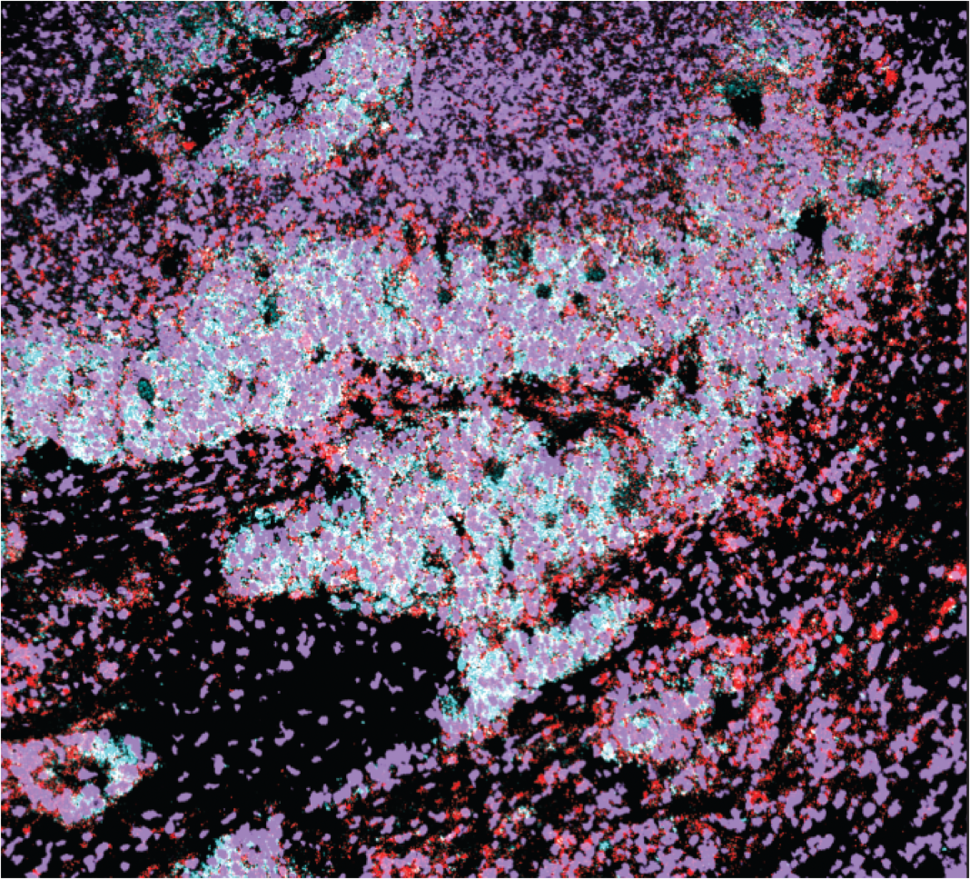
CMR: We used a very informative imaging technique called multiplexed ion beam imaging (MIBI). The single-cell profiling we used does not capture spatial information; you have no idea which cell talks to the other cell and you have no idea what the structure of the tumor or what the local environment looks like.
With MIBI, you get a map of what cells are in adjacency and which cells might be communicating within local environments. MIBI allowed us to confirm a lot of our scRNA-seq observations and also provided a lot more insights into how the different cells might be interacting within the tumor and how the tumors are interacting with the immune system.
PIW: We’ve discussed a lot about the technical feat that is single-cell atlas building. Can you provide some background on SCLC? What exactly are “small cells” and what is their normal function?
CMR: Small cells are cells that have a neuroendocrine phenotype, i.e., they express some proteins that we see in endocrine tissues and in neuronal tissues. In normal development, there are neuroendocrine precursors that are scattered all over your body. The cells are also small, of course. Under the microscope they have almost no cytoplasm and you see sheets of really tiny cells.
One theory holds that SCLCs come from neuroendocrine precursors which get mutated by exposure to tobacco carcinogens. Alternatively, we know that SCLC can arise from a normal airway cell that lines the air sacs of the lung and that usually gives rise to lung adenocarcinoma, if it acquires the same mutations that create SCLC.
PIW: What was known about SCLC prior to building a single-cell atlas?
CMR: SCLC is a particularly lethal form of lung cancer and we've made very little progress in terms of improving clinical outcome over the past 30 or 40 years. It's about 15% of all lung cancers. Most patients are metastatic at the time of diagnosis and have a median duration of survival of only about 13 months.
We knew some of the key driver mutations in SCLC and the overall genomic landscape. It is a tumor type where we often see loss of key tumor suppressor genes rather than gain of activating mutations (which is common in other kinds of lung cancer). The loss-of-function as opposed to gain-of-function alterations are very difficult to target therapeutically.
PIW: What are the major findings you discovered about SCLC?
CMR: Perhaps the most interesting and surprising thing is we found a cell state within SCLC that had not been previously described—a state with very stem-like characteristics that is associated with poor outcome in patients. It is a population of cells that highly expresses a gene called phospholipase C gamma 2 (PLCG2). We think it is maybe a primitive stem-like cell that may give rise to other cell types within the tumor.
I’ll note that these PLCG2-high cells are not different from the other cells within the tumor in terms of the mutations they have. But functionally they are very different!
PIW: So this PLCG2-high state (and really any of the cell states) can occur in each of the different subtypes of SCLC?
CMR: Yes! So previously, we’ve defined SCLC tumor types according to which of three transcription factors is highly expressed: ASCL1 (A), NEUROD1 (N), and POU2F3 (P).
Regardless of these tumor types (or the patient’s treatment history, or tissue site), we found a group of cells with upregulated PLCG2, along with pathways related to metastasis and neural stem cells. This population of cells recurs across these otherwise very different SCLC samples and predicts worse survival, so we’re very interested in figuring out what these cells are really doing!
Another interesting thing we found was that some cells seem to be an intermediate SCLC subtype: between A and N or between A and P. We knew that these types are interconvertible—cells can go from one to the other and tumors as a whole can go from predominantly A to N, for example. But from the single-cell data, we could see cells that were perhaps in the process of flipping from one state to another.
This points to the plasticity of SCLC tumors—their ability to kind of shapeshift and change. This plasticity may also be limiting our ability to effectively treat these tumors.
PIW: Can you speak to how quickly or often this switching might be happening in any given patient? Are the cells going back and forth between states or just in one direction?
CMR: We don't really know the timing yet. What we do know is when we treat SCLC patients, we often get great responses to chemotherapy initially. We get tremendous shrinkage of the tumor and yet the tumor comes back chemo-resistant and lethal in a median to 13 months. So something shifts in the course of those few months. And that's something we're really interested in trying to understand and model in the lab.
PIW: Did the single-cell analysis provide insights into what is happening with the immune system in SCLC?
CMR: SCLC has been relatively resistant to immunotherapy, a type of therapy that has transformed how we treat some other cancers. Even though first line therapy for SCLC does now include immunotherapy, only a small fraction of the patients respond durably. So we're really trying to study why these tumors don’t respond to immunotherapy and what can we do to make them more immunoresponsive.
These cancers usually have many mutations, which means that the immune system should have a lot of substrate to recognize these tumors as foreign and fight the tumor. And yet, they're typically not very immunoreactive. So we’d like to better understand and try to reverse that.
In our study using MIBI and flow cytometry data, we found few immune cells infiltrating the tumors, less than what is found in lung adenocarcinoma. In samples that did contain immune cells, they were compartmentalized rather than intermixed with SCLC cells.
PIW: Is there a piece of information or type of data you wish you could have about SCLC to understand it better?
DP: Clinical data is extremely heterogeneous compared to cancer models, so having greater numbers of samples can always help.
PIW: What biological question are you really interested in answering?
DP: We are really interested in understanding how genetic mutations endow tumor cells with plasticity, which we discussed earlier. Plasticity endows tumor cells with adaptability (without need for further mutation), enabling the cells to evade the immune system, resist therapy and metastasize to new environments. If we can understand the the underlying mechanism, we can directly curtail this plasticity, making the tumor more vulnerable to treatment.
CMR: If we can understand what drives cells from one state to another, can we “trap” the cells in a particular state, perhaps making the tumor more treatable?
And as I mentioned before, what is this PLGC2-high population really doing? I think it may be a key to the ability of this tumor to self-regenerate. These are two big knowledge gaps that may or may not be related. But I think we really need to think hard about the biology and think about strategies to attack this tumor.