Genomic underpinnings of brain tumors expanded
- Posted:
240-760-6600
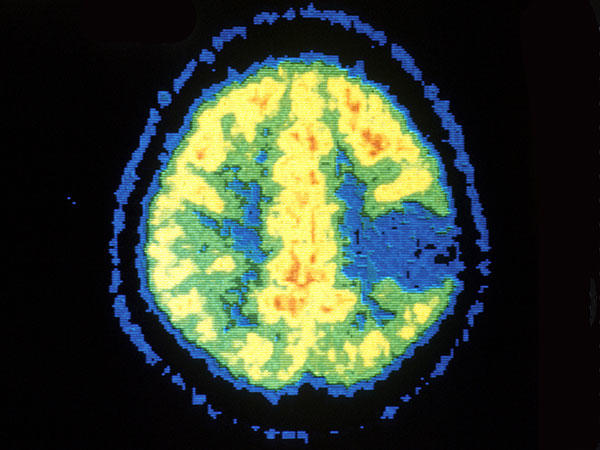
PET scan showing astrocytoma.
Credit: National Cancer Institute
Glioblastoma multiforme (GBM), the highest grade and most-aggressive form of glioma, a type of human brain tumor, was the first cancer type to be systematically analyzed by The Cancer Genome Atlas Research (TCGA) Network, led by the National Cancer Institute (NCI) and the National Human Genome Research Institute, both parts of the National Institutes of Health. Initial results of this analysis, including the identification of significantly mutated genes, were reported in 2008. In 2013, TCGA’s analysis of GBM was expanded to identify several additional significantly mutated genes. In a new study, which appeared June 10, 2015, in the New England Journal of Medicine, TCGA researchers analyzed nearly 300 cases of diffuse low- and intermediate-grade gliomas, which together comprise lower-grade gliomas (LGG). LGGs occur mainly in adults and include astrocytomas, oligodendrogliomas and oligoastrocytomas. Due to their highly invasive and diffuse nature, complete surgical removal is nearly impossible, and the residual tumor can result in spread of the cancer or disease progression, but at differing rates. Unlike GBMs, these tumors have highly variable clinical behaviors, with a subset of LGGs progressing to GBMs within months, whereas others remain stable for years. Although these clinical differences are not easily predicted by how the tumor cells appear under a microscope, the potential relationship between the molecular features of these tumors and their clinical behavior has not been explored comprehensively.
In LGGs, TCGA researchers found recurrent mutations and chromosomal copy number changes that were associated with clinical outcome. The researchers honed in on a specific chromosomal abnormality called 1p19q co-deletion, which is characterized by the simultaneous deletion of the short arm of chromosome 1 and the long arm of chromosome 19. This abnormality has been linked to a more favorable clinical outcome. Tumors that had 1p19q co-deletion also frequently had mutation of the IDH1 and IDH2 genes, and this combination was associated with the most favorable clinical outcomes. The large majority of patients without IDH mutations showed strikingly similar genomic aberrations to GBM and had a similarly inferior outcome. These findings suggest that this subtype of LGGs may be the precursor to some GBMs. Current research is aimed at developing drugs to inhibit mutant forms of the IDH1 and IDH2 proteins, which might provide benefit to patients with the less-aggressive molecular subtype of LGG. Conversely, because LGGs without IDH mutations are similar to GBMs both molecularly and clinically, patients with such tumors may benefit from treatment protocols being used for GBM patients. According to Daniel J. Brat, M.D., Ph.D., Emory University Hospital, Atlanta, corresponding author of the LGG study, this research should help redefine relevant classification of LGGs to incorporate molecular data with the pathologic diagnosis so that patients can receive the most optimal therapy available.
###