RAS Initiative Bioinformatics Research
The RAS Initiative Bioinformatics Research Team provides data processing and analytical support through three broad approaches:
- Processing and analysis of data from RAS Initiative colleagues or our collaborators (See example publication.)
- Exploratory or hypothesis driven data mining for RAS Initiative colleagues (See example publication.)
- Data mining of public data (e.g., TCGA, DepMap) for novel insights of RAS biology (See example publication.)
We also created software tools to organize and interpret data produced by the RAS Initiative. We imported “big data” from public data repositories and analyzed them with a focus on the genes in the RAS pathway. By integrating internal and external data, we help improve understanding of RAS-driven cancers.
Progress
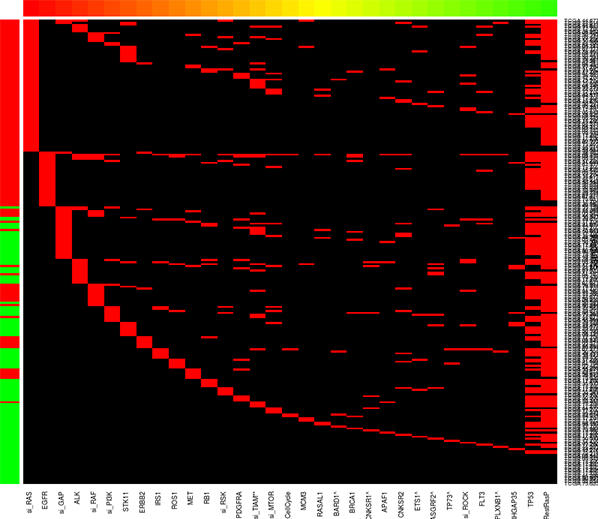
The RAS Initiative, together with groups from the University of California, San Francisco, and the Broad Institute in Boston, have probed dozens of cancer cell lines for the effect of knocking down entire signaling nodes (such as all three RAS genes, all three RAF genes, etc.). Learn more about this project.
The RAS Bioinformatics Research Team is integrally involved in drug discovery projects targeting KRAS and RAC1 and is responsible for processing raw data and analysis, data visualization and side-by-side comparison of selected mutant targets, and curve fitting for dosage data of selected candidate lead compounds.
By data mining of public expression and survival outcome data from CCLE and TCGA as well as internal siREN data, we derived computational RAS-dependency indexes (RDIs) as putative measures of RAS dependency. Our RDIs correlate well with experimental data. The computational RDIs were expanded to cancer patients and have potential clinical applications. Learn more about this study.
Our team has analyzed the “big data” produced by The Cancer Genome Atlas in the context of mutant or wild-type RAS genes and found intriguing biological connections between mutation status and their expression levels, published here.
Last year, we helped update a NCI Dialogue describing co- and contra-mutation patterns with KRAS using AACR’s GENIE, BROAD Institute’s GDAC and cBioPortal databases with cancer patient samples for KRAS tumor types including PAAD, LUAD, COAD, and READ.
Besides data mining and informatics support, we developed in-house tools, such as a survival analysis method that can help avoid a common pitfall of survival analysis. Learn more about this in-house method.
Finally, we have built an architecture that stores, organizes, searches, and retrieves the data produced within the RAS Initiative.
Projects
By providing sophisticated bioinformatic tools, we free the biologists from tedious data processing and complicated data analysis so they can focus on the wet-lab experiments. Our informatics support guides the interpretation of data and often provides direct insights into the underlying biology. Furthermore, our data mining efforts yield novel hypotheses in RAS biology that benefit the ongoing internal projects as well as the RAS research community.
- Analysis of siREN data (using pools of siRNAs to knock down signaling nodes in cancer cell lines) and integrating it with drug sensitivity data from the Genomics of Drug Sensitivity in Cancer, which was published in Cell Reports and available on PubMed.
- Processing and analysis of the ongoing internal drug screening data
- Support and implement data mining projects with our colleagues from RAS program and NCI. (Learn more from the NRF2 pancreas project and the RAS vs metabolic dysregulation project)
- Analysis of RAS and RAS pathway gene expression in tumor and normal samples in The Cancer Genome Atlas (TCGA) data, published here
- Data mining of public expression and survival outcome data from CCLE and TCGA as well as internal siREN data (Learn more from the published study.)
- Ongoing data mining of expression, mutation, and CRISPR screening data from DepMap
- Development of tools to store, track, and retrieve data from FNLCR RAS Initiative projects
Tools
The tools that help us fulfill our mission are mainly derived from public assets. These public or access-controlled cancer related databases and portals provide us the data resources for our data mining projects.
The servers and databases maintained by our organization provided us with the computational and analytical power for data processing and data analysis.
- Programming, web, and scripting languages
- Public genomics and cancer data sets such as IGCG, TCGA, GENIE, and PCAWG
- Databases including Oracle, mysql, and Filemaker
- Cell line datasets such as nci60, CCLE, DepMap, COSMIC, and GDSC
- Data portals such as Biomart, cBIOportal, and caHUB
- In-house developed pathway analysis method: PPEP, survival analysis method: GradientScanSurv
The RAS Initiative Informatics Research Team
Team lead and contact
Dr. Ming Yi
yiming@mail.nih.gov
301-846-5764
Collaborators
- University of California, San Francisco
- Department of Statistics, University of Illinois Urbana-Champaign
- Broad Institute in Boston