Introduction
Broad sharing of biomedical research data is essential to not only further our mechanistic understanding of disease, but also to enable swift development of safe and effective therapeutic interventions.
Data sharing facilitates transparency in science, allows for secondary validation of findings, and accelerates the generation of novel hypotheses and discoveries, compounding the impact of the original study.
In 2003, NIH set expectations that data generated through support of NIH grants, cooperative agreements, and contracts be made widely and freely available while protecting research participants. The expectation was further iterated within the NIH Genomic Data Sharing Policy of 2014, which required sharing of large-scale sequencing studies.
The new NIH Data Management and Sharing (DMS) Policy, that went into effect in January 2023, requires prospective planning of the management, preservation, and timely sharing of biomedical data for all NIH-funded research projects. The National Cancer Institute (NCI) is committed to this policy and aims to shift the culture of cancer research towards consistent sharing of high-value data, while acknowledging and addressing the data sharing challenges currently faced by the scientific community.
The NCI Division of Cancer Biology (DCB) supports basic cancer biology research, which generates valuable insight into the fundamental mechanisms of cancer initiation, progression, and response to therapy. Most research awards (70%) supported by DCB are investigator-initiated research project grants (R01).
Traditionally, cancer biology-focused R01 projects generate data to address a specific hypothesis and, mostly do not support large, cohort-based sequencing studies. Multiple studies have quantified the value of sharing data, but the impact of sharing cancer biology datasets generated through the support of traditional R01s is less clear.
To capture the value of sharing and reusing fundamental cancer biology data, here we quantify the impact of broadly shared and reused datasets published in 2018-2020 that were generated with support of DCB-managed R01 awards.
We show that publications that shared cancer biology datasets were more highly cited and were within higher cited research fields. Additionally, publications that reused these datasets had higher citation rates and influence on their specific research fields, demonstrating the direct benefit to researchers in sharing and reusing small- to medium-sized cancer biology datasets.
The 2003 NIH Data Sharing Policy set the expectation of broad data sharing, many cancer biology investigators have only recently been required to prospectively plan for broad sharing at the time of publication.
Using data collected from the basic cancer research community via an NIH RFI, we comment here on the understanding, concerns, and current challenges for NIH DMS plan development, management, and implementation. The RFI results also highlight the importance of establishing field-wide standards for data sharing and management.
Publicly Shared Cancer Biology Datasets Are Mainly Composed of Genomics Data
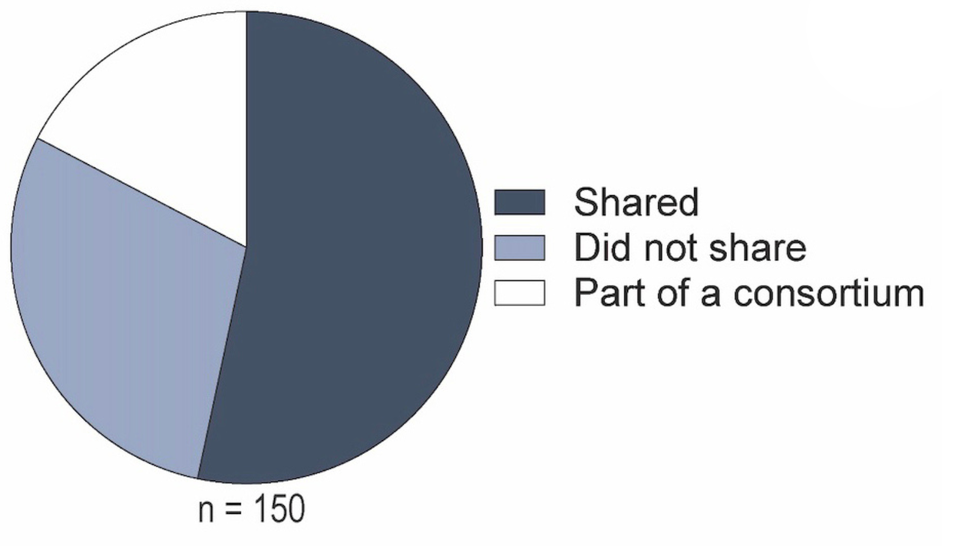
To investigate the impact of sharing cancer biology datasets, we assessed the data sharing status of DCB R01-funded publications between 2018 and 2020. We focused on the 150 most influential publications, as measured via Relative Citation Ratio (RCR), or the field- and time-normalized citation rate.
The median RCR for an NIH-funded publication each year is 1.0. Thus, an RCR of 2.0 implies that a publication received twice as many citations as the median NIH-funded publication in its co-citation network.
The 150 publications analyzed were ranked in the top 1.1% of highest cited publications in their respective co-citation networks. Across these publications, we identified those that shared trackable dataset identifier(s), including accession codes, digital object identifiers, and links to repository depositions. These data were often, but not always, provided within data availability statements.
- 53% of the publications shared a trackable dataset
- ~ 17% of the publications were authored by a research consortium (e.g, TCGA, GTEx, and PCAWG)
Given the highly collaborative nature and additional data sharing requirements of investigators within these consortiums, the associated publications were removed from further analyses. This allowed for investigation into the sharing of data generated from small- to medium-scale research endeavors, which we equate to those studies supported by NIH R01 awards.
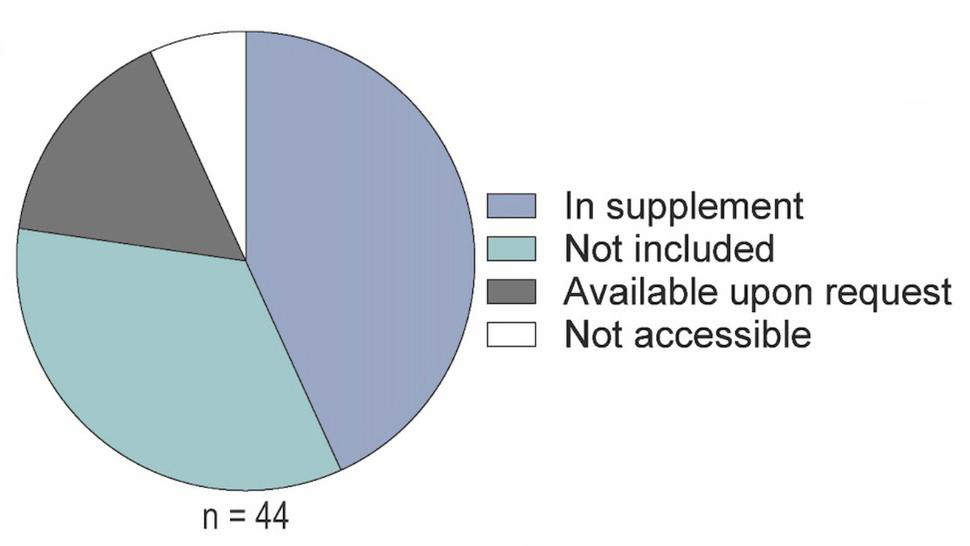
Of the publications that did not share trackable dataset(s):
- 43% included additional data within supplementary materials
- 16% stated that data were available upon request
- 7% provided shared data that are no longer available
- 34% neither shared data nor provided a data availability statement
- 90% of the 150 publications addressed data sharing to varying extents
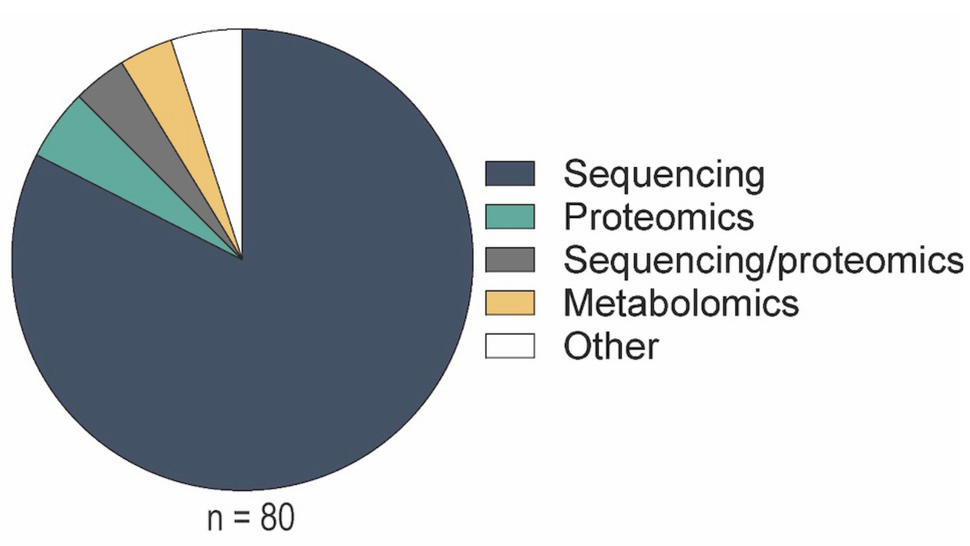
Of the publications that shared trackable dataset(s):
- 82.5% shared sequencing datasets
- 17.5% shared proteomics, metabolomics, and other data types (e.g., ligand docking, microarray, and clinical data)
High rates of sharing sequencing datasets may be a result of the implementation of the NIH Genomic Data Sharing (GDS) policy in 2015, which set expectations to ensure the broad and responsible sharing of genomic research data.
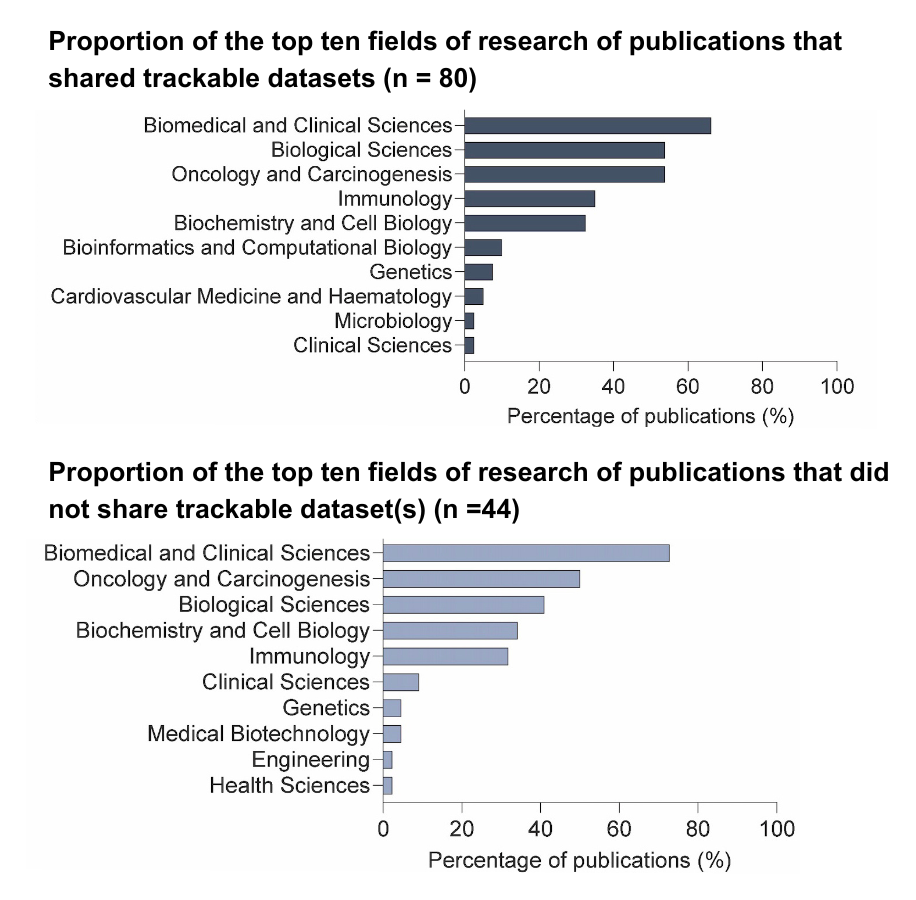
To delineate the types of research studies that are generating and sharing dataset(s), we utilized a commercial language model, Dimensions AI, to assign each publication to fields of research based on the Australian and New Zealand Standard Research Classification.
There were similarities in the research fields of publications that did and did not share data, encompassing biomedical and clinical sciences, biological sciences, oncology, immunology, and genetics.
Overall, data sharing of predominantly genomics data spanned across several relevant cancer biology fields; however, 30% of studies in these fields did not share trackable datasets.
Publications that Shared Datasets had Higher Citation Rates and Were Published in More Highly Cited Fields
We next juxtaposed the bibliometrics of publications that did and did not share trackable dataset(s) to determine if sharing trackable datasets impacts the influence of publications within the research community.
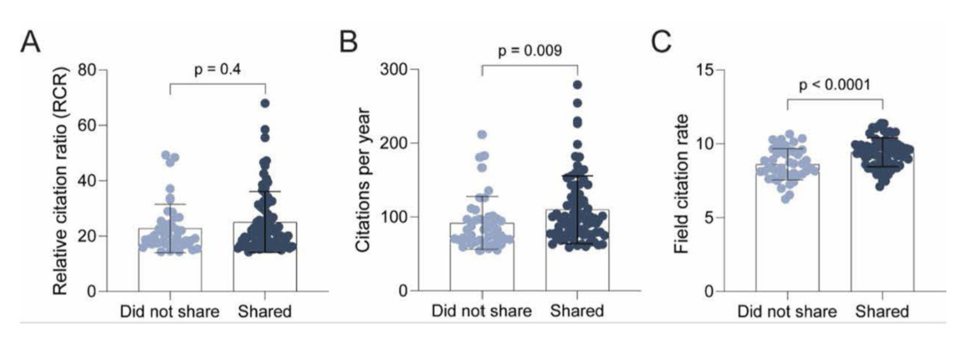
We found that there were no significant differences in RCR between publications that shared and did not share dataset(s). However, as this analysis focused on DCB-funded publications with the top ~1% highest RCR scores, differences in publication influence might be difficult to detect because all the publications analyzed are highly influential (as measured by RCR).
We next extracted the citation rate of publications to directly investigate the effect of data sharing on publication citations. We found that publications sharing trackable dataset(s) had a 20% increase in citations per year compared to publications that did not share trackable dataset(s). Since this increase was not evident in the RCR, it suggests that there were also shifts in the field citation rates.
We further probed this aspect and found that the average field citation rate of data-sharing publications was 9% higher than the field citation rate of publications that did not share data.
Therefore, while publications that shared data were cited more frequently, these publications were published in fields that were overall more highly cited by the research community, suggesting that data sharing is not the sole driver of the increase in citations of these publications but that a relationship exists between highly cited fields and the propensity to share data.
Publications that Reused Cancer Biology Dataset(s) Had Higher Influence
We next tracked reuse of datasets shared by DCB R01-funded publications described above. Using
Google Scholar, we searched for dataset identifier(s) and confirmed datasets were used in research publications.
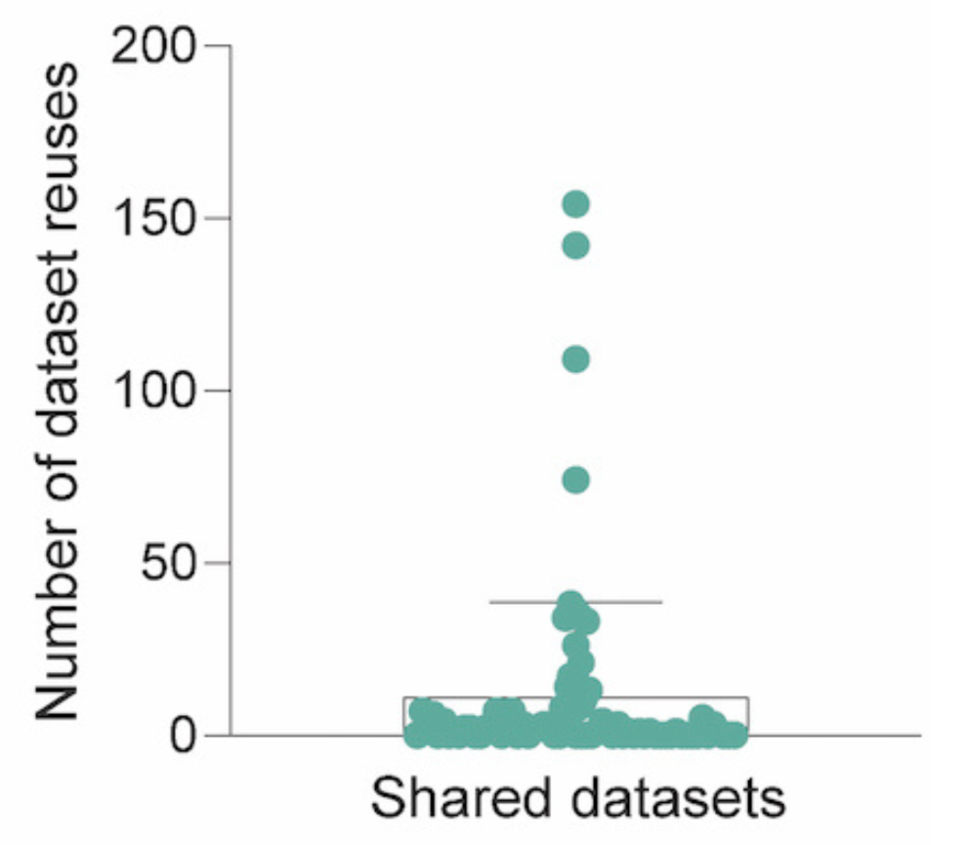
Of the shared dataset(s):
- >60% were reused in at least one subsequent publication
- 11 identified reuses on average per dataset
- 89% of reuses were in open-access publications
After identifying publications that reused the shared dataset(s), we investigated the characteristics of secondary use. First, we wanted to confirm that the authors in the original dataset generation were not involved in secondary data reuse publications, and found that 93% of secondary data reuse was by research teams completely unique to the team generating the original dataset.
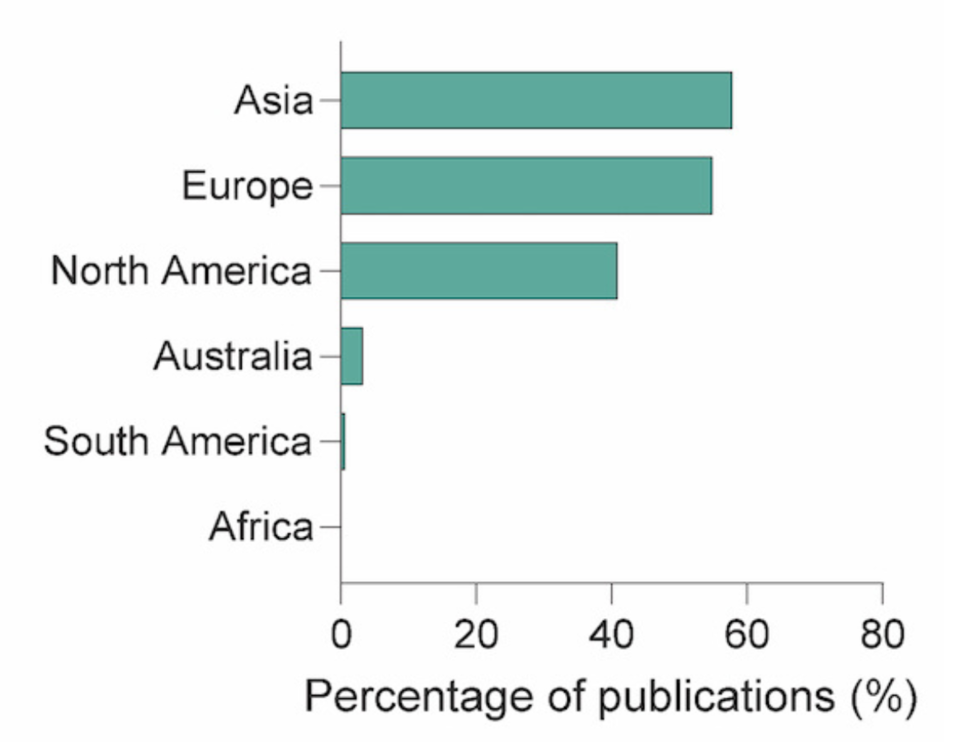
Additionally, Web of Science was used to examine the geographical locations of the primary institutions where publications that reused the 80 shared trackable datasets originated:
- 58% - Asia
- 55% - Europe
- 40% - United States
This demonstrates the global reach of these data.
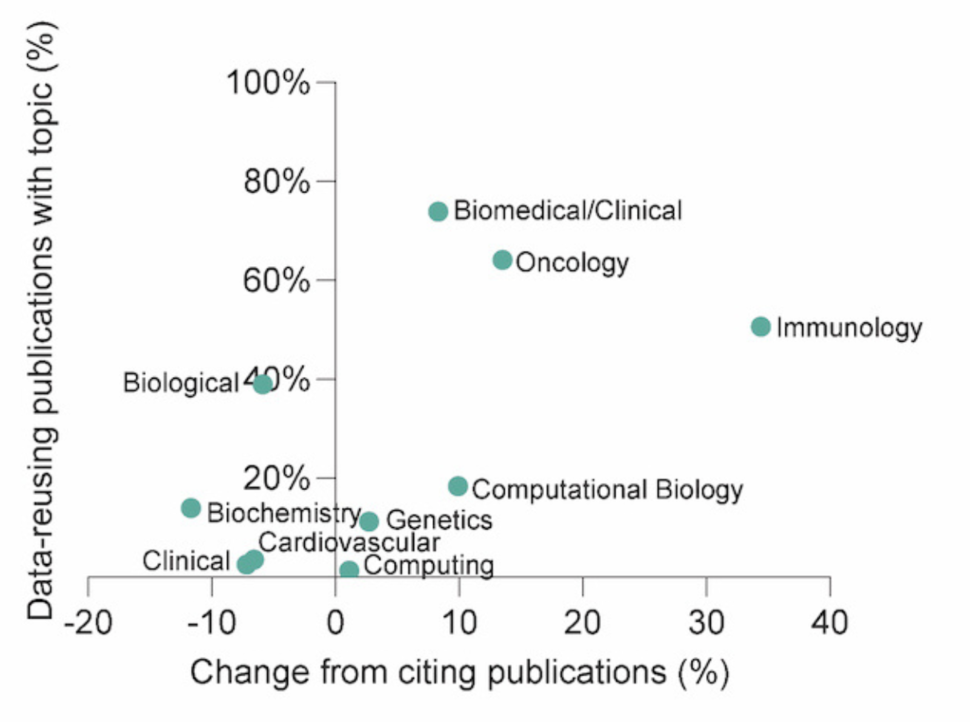
Using Dimensions AI again, we assessed the most prominent research areas of the publications reusing data to understand which sub-fields are employing these datasets for secondary use.
When compared to publications that only cited the data sharing publications without reusing a dataset, there was an increased proportion of studies in the following areas:
- Immunology (34%)
- Oncology and carcinogenesis (14%)
- Bioinformatics and computational biology (10%)
- Biomedical and clinical sciences (8%)
These are research areas that typically utilize larger datasets and thus may be more equipped for secondary reuse.
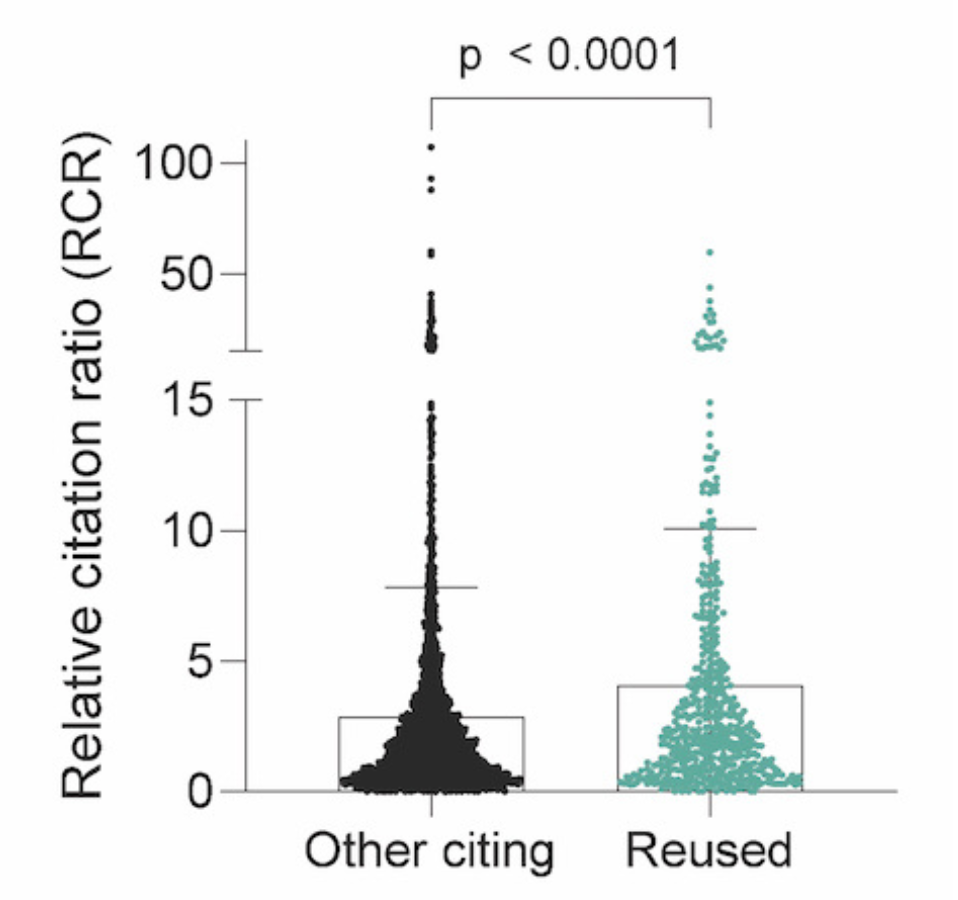
Lastly, we quantified the effect of reusing publicly shared cancer biology datasets on the publication influence. We found publications that reused dataset(s) had a 41% increase in RCR compared to publications that only cited the original data-sharing publications.
This indicates that publications that reused dataset(s) had more influence in the research community and received more citations than publications that only cited the original publications.
Thus, these shared cancer biology datasets facilitated reuse of these data by numerous research fields globally and ultimately contributed significantly towards more influential research studies.
Dataset Reuse Enabled Data Validation, Provided Insight into Biological Mechanisms, and Supported the Development of Computational Tools
We next asked how DCB-supported data sets were reused. This analysis was focused on five publications with widely used dataset(s) and categorized how datasets were employed in subsequent studies. The five publications shared different types of genomic sequencing data.
Case Study Publications for Analysis of Dataset Reuse
| Publication (PMID) | Dataset identifier(s) | Shared dataset type |
|---|---|---|
| Azizi et al. Cell 2018 (2996157) | GSE114725 & GSE114727 | Single-cell RNA-seq of human breast tumor immune cells (CD45+) |
| Chapuy et al. Nature Med 2018 (29713087) | phs000450 & GSE98588 | Gene expression profiling of human diffuse large B-cell lymphoma (DLBCL) tumors |
| Elyada et al. Cancer Discov 2019 (31197017) | GSE129455 | Single-cell RNA-seq from pancreatic ductal adenocarcinoma mouse (fibroblast-enriched population) |
| Huang et al. Nat Cell Bio 2018 (29476152) | GSE90639, GSE90642 & GSE90684 | RNA immunoprecipitation-seq of HEK293T cells with Flag-tagged IGFBP1/2/3/ overexpression, RNA-seq of IGF2BP knockdown and control HepG2 cells, & N6-methyladenosine (m6A)-seq of METTL14 knockdown HepG2 cells |
| Sade-Feldman et. al. Cell 2018 (30388456) | phs001680.v1.p1 & GSE120575 | Single-cell RNA-seq of immune cells from human melanoma patients treated with checkpoint inhibitors with clinical outcomes |
Dataset(s) shared in these publications were reused between 26-154 times.
- 82% of the publications that reused these dataset(s) cited the original publication
- 18% shared only the associated dataset identifiers without providing direct credit to the original publication.
Through manual curation of publications, we identified six main types of dataset reuses:
- Support of a biological claim
- Validation of a biological finding
- Insight into a biological mechanism
- Development of a computational model
- Validation of a computational model
- Creation of a data resource
The ways in which the shared dataset(s) were reused was highly dependent on the context of the original dataset(s) generation and the data type.
For example, publications that reused data from Huang et al. in which the data were originally generated to reveal mechanisms in stabilizing m6A-modified mRNAs, mainly did so to delineate additional new biological mechanisms.
Publications utilizing data from Sade-Feldman et al., which generated single-cell RNA-seq of immune cells in tumors of patients with melanoma treated with checkpoint inhibitors and provided the associated clinical outcomes, mostly used the data to support the clinical relevance of their biological findings. In addition, this dataset also had the highest data usage for the purpose of developing a computational model, such as a predictive model of immune checkpoint therapy response.
Overall, the datasets were reused for many different purposes, mostly by unique investigators to develop and support new research findings in the cancer biology community. This demonstrates that the impact of sharing data can go far beyond that of its original intention.
Opportunities to Support Improved Data Sharing by Investigators
To better understand the obstacles in sharing cancer biology data, NCI solicited feedback in the summer of 2024 on the experiences of cancer biology investigators in developing and implementing data management and sharing plans along with sharing and reusing datasets. "Request for Information (RFI): Data Management and Sharing of Cancer Biology Research (NOT-CA-24-062)" received responses from researchers at different career stages, including postdoctoral fellows, tenure- and non-tenure-track faculty, and research administrators.
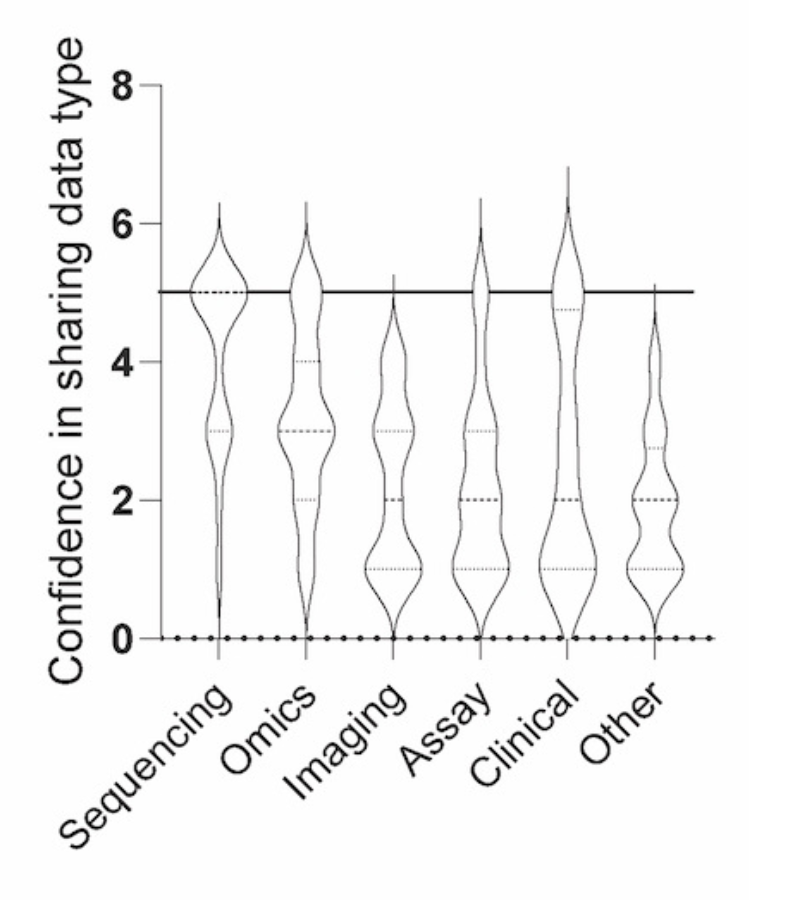
We first sought to compile the confidence level in sharing various data types associated with cancer biology research. Investigators were most confident in sharing sequencing and omics datasets, while lacking the same confidence in sharing imaging, molecular and cellular assay, clinical, and other data types. Encouragingly, when asked about confidence in reusing shared datasets, respondents were moderately confident (scoring 3.7 ± 1.1 out of 5).
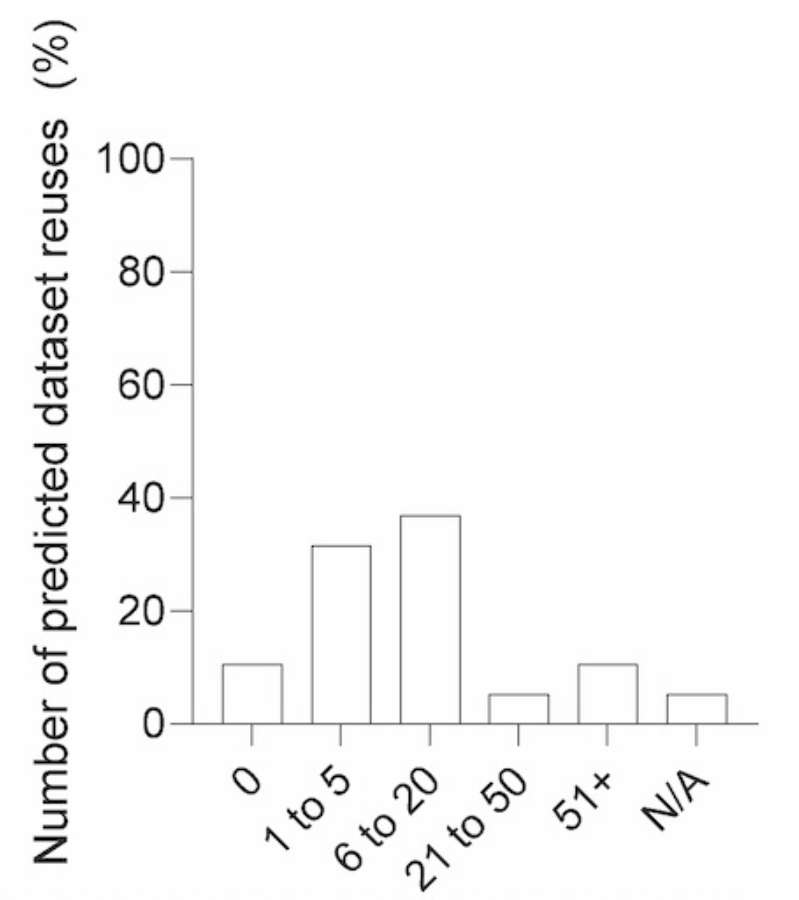
We next assessed how confidence in reusing datasets related to the perceived impact of sharing cancer biology datasets:
- 53% responded that sharing their cancer biology data would lead to more than 5 secondary uses
- 53% responded that sharing of cancer biology data has led to an increase in collaboration with other investigators (demonstrating that data sharing has already benefited respondents in this small sample size)
Despite the perceived benefits by cancer biology researchers, 29% of the publications analyzed in this study did not share datasets.
While sharing is now required under the NIH DMS Policy, we recognize that obstacles still exist for investigators in developing and implementing plans to share data.
To explore these obstacles further, we examined the most time-intensive components of the NIH DMS Policy.
The components of writing DMS plans that took the most time included:
- Identifying data standards
- Determining proper data repositories
The most time-intensive components of implementing DMS plans included:
- Formatting data/metadata
- Collecting metadata
Less than half of the respondents were aware if their institution provided training on data management and sharing, and an equal number felt properly trained to develop and share publicly available datasets.
Thus, increasing awareness of field-specific standards on data collection and formatting of data and metadata, along with providing institutional training on data management and sharing, could alleviate some of the administrative burden cancer biology researchers experience in broad sharing of FAIR (findable, accessible, interoperable, and reusable) datasets.
Discussion
Broad sharing of cancer biology data accelerate hypothesis and discovery driven research. It also extends the reach of cancer biology studies thereby enhancing NIH’s mission to seek fundamental knowledge of living systems and applying this knowledge to enhance public health.
However, there remain concerns amongst researchers that hinder the extent to which they share datasets. According to a 2022 survey of more than 6,000 researchers, the top motivating factors to share data include expectations set by the research field and/or requirements set by publishers and funding agencies.
The NIH DMS Policy, effective since 2023, sets the requirement for prospective planning of the management, preservation, and timely sharing of biomedical data for all NIH-funded research projects. Such policy changes help shift data sharing culture, as was seen by the high frequency of genomic data sharing by DCB-supported publications following the implementation of the NIH Genomics Data Sharing Policy.
While prior studies have shown increased citations due to data sharing, we specifically aimed to quantify the bibliometrics associated with sharing of small- to medium-scale basic cancer biology data. We show publications that shared cancer biology datasets had a 20% increase in citation rate compared to those that did not share datasets in their publications. However, the overall impact of the former was not different than other publications in their co-citation network.
Interestingly, our data also suggested a relationship between the degree of data sharing and the overall field citation rate. A larger study including publications across a broader range of RCR values is needed to fully delineate the direct impact of data sharing on field citation rate.
In this study, we found using Google Scholar facilitated discovery to a greater extent than PubMed or other search methods. However, the discovery of datasets using data object identifiers or accession numbers required significant manual curation. Analyzing data sharing status of larger collections of publications will require large language models to help with data curation.
Our analysis demonstrates that secondary data reuse has a significant positive influence on publication impact, independent of research field. Several factors might contribute to this result, including an increased confidence in study conclusions that are supported by independently generated datasets.
These data illustrate how the impact of shared data is propagated within the research community and that investigators within specific fields of research, such as immunology, oncology, and computational biology already place an increased value on studies that reuse shared data.
Our study provides an exciting preview of how all data types generated within R01-supported projects, shared under the new NIH DMS policy, will have a lasting impact on the research community.
Despite the positive impacts of sharing and reusing datasets, there are many opportunities to further improve broad data sharing. Some approaches include:
- Supporting data sharing and management training
- Establishing data sharing standards across various data types generated in cancer biology
To address some of these gaps in data sharing practices, DCB has provided guidance for the NIH DMS Policy for investigators performing basic cancer biology research. Along with providing DMS guidance for putting together a good DMS plan, it offers requirements, resources, standards, and additional policy information, which will help investigators prospectively plan their data management and sharing.
There are additional opportunities for the research community to collaborate and establish workflows that can be adopted by the field to generate FAIR data and train future generations to readily integrate these workflows into their research, ultimately reducing the administrative load of addressing these requirements.
In summary, we show that the sharing of basic cancer biology data is associated with an increase in citation rate and led to an average of 11 secondary uses of the datasets. These datasets were reused by cancer researchers around the world and supported or validated new findings or computational models.
Adoption of the NIH DMS Policy data sharing practices and receiving DMS plan writing resources and training and will empower researchers to create more FAIR datasets and amplify the impact of their research.
Materials and Methods
Approaches related to this study, including the identification of data sharing in cancer biology studies, bibliometric analysis, the analysis of data reuse, and statistical analysis, can be found in a Materials and Methods document.