Can the New “Omics” on the Block Find Liver Cancer in Blood?
, by Nadia Jaber
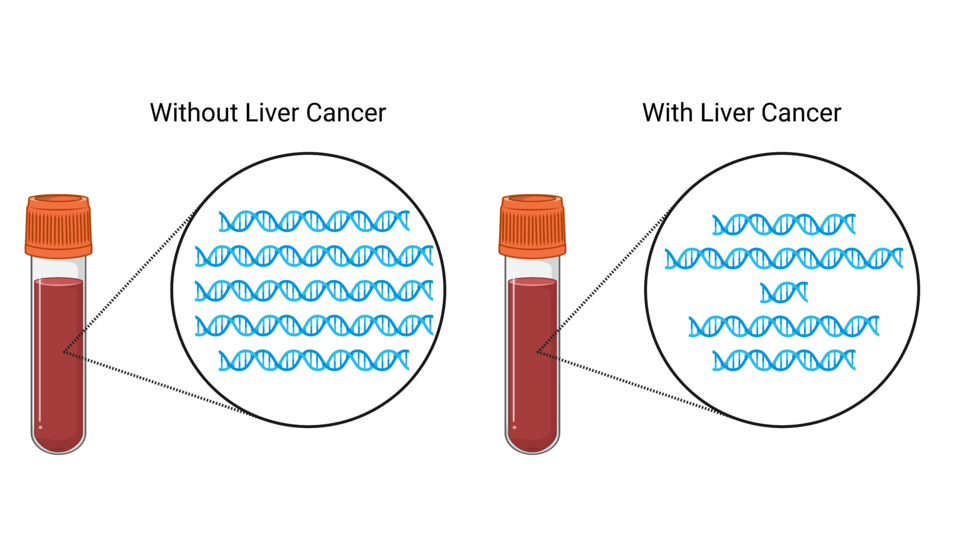
Researchers have developed a blood test that, in a preliminary study, accurately detected liver cancer, including in people with early stages of the disease. Unlike most other blood tests in development for cancer detection, this one uses a new type of technology, called fragmentomics, to analyze bits of DNA in the blood.
Liver cancer is the third leading cause of cancer death worldwide. When the disease is caught early, upwards of 70% of people survive more than 5 years. But if it’s found after the cancer has spread, that number drops below 20%.
For that reason, people who have a high risk of liver cancer—such as those with cirrhosis or chronic hepatitis B virus infection—are advised to get regularly monitored for the disease. But current tests for liver cancer don’t work very well, aren’t readily available to all who need them, and are expensive.
The new blood test could potentially address those issues, said study co-leader Amy Kim, Ph.D., of Johns Hopkins University School of Medicine.
In the new study, the researchers used machine learning to analyze fragmentomics data and find differences between people with and without liver cancer. When applied to blood samples from hundreds of people—including some known to have liver cancer—the test accurately identified those with liver cancer, including early-stage disease.
They also validated the accuracy of the test using blood samples from another large group of people. The NCI-funded study was published March 1 in Cancer Discovery.
A caveat is the relatively small number of people with liver cancer in the study, Christian Rolfo, M.D., Ph.D., of Tisch Cancer Institute, and Alessandro Russo, M.D., Ph.D., of Papardo Hospital in Messina, Italy, pointed out in a commentary on the study.
Although the results seem promising, the next—and very important—step is to see how the blood test performs in a larger group of people who haven’t yet been diagnosed with liver cancer, noted Tim Greten, M.D., a liver cancer expert in NCI’s Center for Cancer Research, who wasn’t involved in the study.
Study co-leader Victor Velculescu, M.D., Ph.D., of Sidney Kimmel Cancer Center at the Johns Hopkins University School of Medicine, and his colleagues are planning follow-up studies of people at high-risk of liver cancer to see if the approach can catch early-stage disease.
Liquid biopsies and fragmentomics
Scientists have long known that both healthy and cancerous cells shed pieces of their DNA into the bloodstream. More recently, researchers have found that these DNA fragments can act like breadcrumbs, leading straight back to the cancer cells that tossed them out.
That revelation has led to a huge surge in the development of blood tests that detect cancer, said Dr. Greten. Such tests are often called liquid biopsies.
Most liquid biopsy tests developed to date rely on genomics (scanning DNA in blood for cancerous mutations) or epigenomics (analyzing the pattern of chemical tags on DNA in blood). Fragmentomics, in contrast, looks at the pattern of the amount and sizes of DNA fragments in the blood.
In 2019, Dr. Velculescu and his colleagues developed a fragmentomics approach called DELFI that analyzed DNA fragments from throughout the genome.
In a preliminary study, they used DELFI to screen for seven types of cancer, not including liver cancer. DELFI was able to pinpoint whether an individual had cancer and what kind of cancer it was, although its accuracy varied by cancer type.
Given the lack of effective tests for liver cancer, the team decided to see if DELFI could detect liver cancer in the blood. They used DELFI to analyze blood samples from 501 people in the United States and Europe, including 75 people known to have liver cancer and 133 people with liver disease (cirrhosis or hepatitis infection) who were at high risk for liver cancer.
People with liver cancer had DNA fragments that greatly varied in size, whereas people without cancer (including those with liver disease) had DNA fragments that tended to be a consistent size, they found.
Additional experiments revealed that these DNA fragments reflect unique patterns of genetic changes, including changes in chromatin organization, known to occur in liver cancer cells.
“These fragments that we’re picking up in cancer patients are not just random. It’s really reflecting the genome of liver cancer [cells],” Dr. Kim emphasized.
DELFI differentiates between people with and without liver cancer
The subtle changes in DNA fragmentation in the blood were enough for DELFI to differentiate between people with and without liver cancer, the researchers found.
And it was very accurate. When separating people with liver cancer and those with liver disease, DELFI scored 0.9 on a 0-to-1 performance scale (also called area under the curve, or AUC). It was even better at separating people with liver cancer from people without liver disease (AUC=0.98). A test with perfect accuracy would have an AUC of 1.
The test also picked up on early-stage liver cancer (AUC=0.9 and 0.81 for the two earliest stages), which can be tricky to spot with other tests.
DELFI was good at finding liver cancer when it was present (high sensitivity) and rarely said someone had cancer when they didn’t (high specificity), the researchers noted.
To validate the test in a separate group of people, the team applied it to blood samples of 223 people from Hong Kong. This group included 90 people known to have early-stage liver cancer and 101 people with known liver disease, mainly hepatitis B infection or cirrhosis.
Again, the test correctly distinguished between people with liver cancer and people with liver disease (AUC=0.97).
Participants in the U.S./Europe and Hong Kong groups had different causes of liver cancer and ethnicities, but the test performed well in both groups, Dr. Kim pointed out. That suggests “that this approach could be generalizable across different high-risk populations worldwide,” Drs. Rolfo and Russo wrote.
Finally, the researchers compared DELFI with a blood test that is used to test for liver cancer in people at high risk of the disease. The test looks for a cancer-related protein called alpha-fetoprotein and is typically combined with an ultrasound of the abdomen.
The alpha-fetoprotein test correctly flagged 52% (39 of 75 people) of those with liver cancer in the US/Europe group, while DELFI caught 85% (64 of 75 people). DELFI also identified a greater percentage of people with early-stage liver cancer than the alpha-fetoprotein test.
Fragmentomics for cancer detection
The research team believes fragmentomics has several advantages when it comes to liquid biopsies for cancer detection.
A challenge with mutation- and methylation-based liquid biopsies is that these cancer-related changes are scarce in the blood, Dr. Kim said, making them hard to find. But with fragmentomics, the DNA doesn’t have to be scanned as intensively and it keeps the cost of the test down, she explained.
Another advantage of fragmentomics is that it requires much less blood than other liquid biopsy tests, she added.
The fragmentomics approach is also appealing because it requires only a blood draw, Dr. Greten noted, which is typically faster, easier to get, and less expensive than an ultrasound.
Fragmentomics is a next-generation liquid biopsy approach, said Dr. Velculescu. And it can potentially be used to detect other kinds of cancer, in addition to those the team has already studied, he added.
A company founded by Dr. Velculescu, and where he serves as CEO, is working on developing DELFI for commercial use.