Dr. Nevan Krogan, a CSBC Investigator and Director of the Quantitative Biosciences Institute at the University of California, San Francisco (UCSF), develops tools to study quantitative changes in biomolecules.
As a graduate student, he studied protein complexes in yeast using mass spectrometry. Now, he is using his knowledge of biological networks to create a map of complex interactions in a cancer cell.
In this interview he discusses mentorship experiences, perspectives on cancer systems biology, and his CSBC research with co-investigator Dr. Trey Ideker.
How do you help trainees accomplish their research and career goals?
My goal is to create an infrastructure and atmosphere that’s conducive for discovery. I want my trainees to work in an environment where they don’t need to worry about the cost of an experiment. Their only limitation should be their imagination. Also, collaborations are key when moving forward in a scientific research career. I help my trainees meet other scientists and push them to network with people at meetings.
Who were your role models during your scientific career journey?
I did my doctorate work at the University of Toronto, where Frederick Banting and Charles Best worked. They were two scientists who discovered insulin in the 1920s and inspired me by what they were able to accomplish in a short period of time.
Within a few years, their basic research results saved millions of lives. More recently, I’m a fan of Steve Elledge. He develops and uses molecular tools to study different types of disease. He has no fear about moving into new research areas and designs new approaches to study systems biology.
What do you think the biggest challenge and opportunity in cancer systems biology research will be in the next 10 years?
The biggest challenge and opportunity are one and the same, which is communication. We, in the field, need to get cancer researchers from different disciplines more effectively talking and working together. A problem in science is that each area is in its own silo. To understand cancer at every level, it’s important to have interactions across scientific fields, including structural biology, chemistry, computational biology, biochemistry, clinical oncology, and more.
Additionally, in science, we often reward the individual, such as the investigator who set up the project or the person who is the last author of a publication. For collaborative research, we need to develop a team-based mentality, and the contributions of the entire team need to be recognized.
Can you describe your CSBC work with the Cancer Cell Map Initiative (CCMI)?
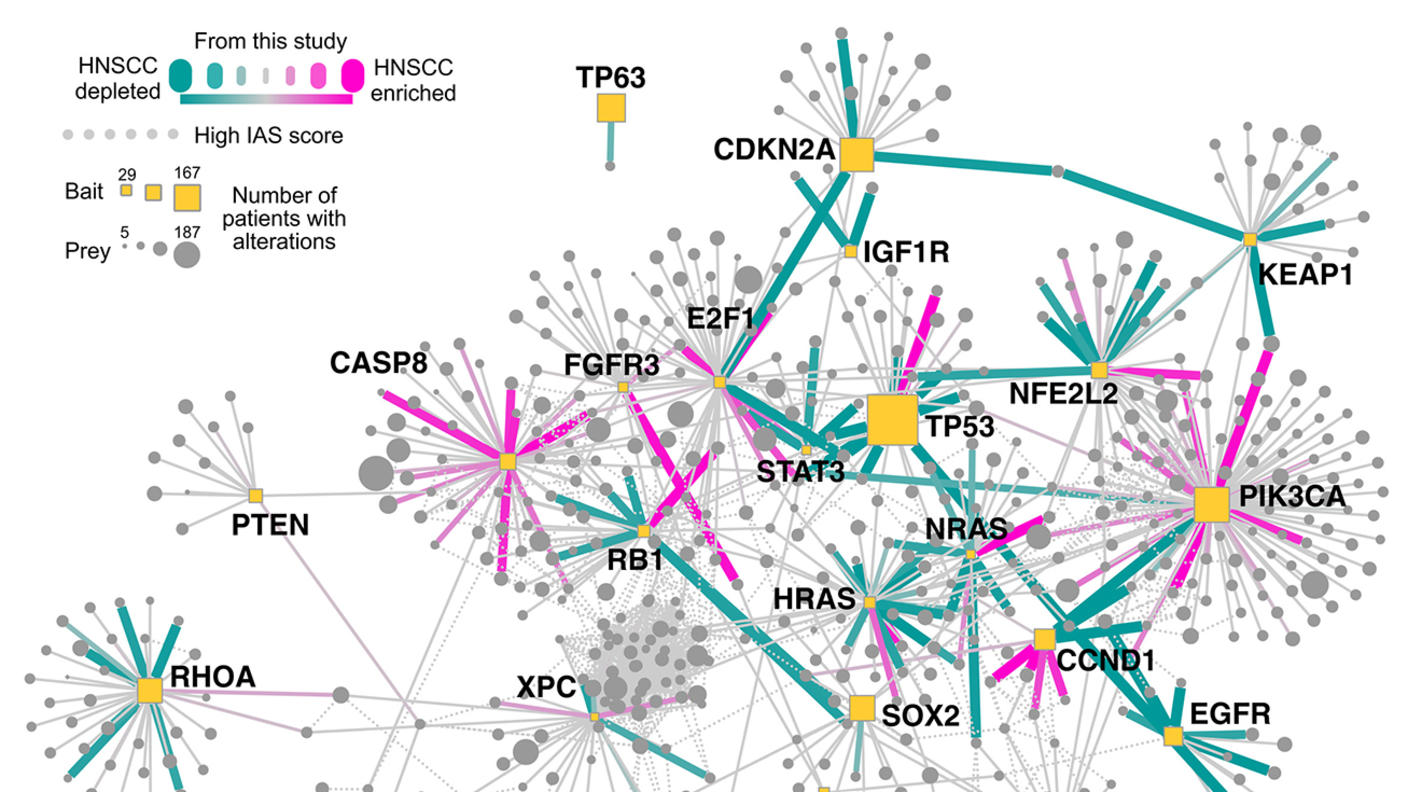
Our CSBC center with the CCMI is unique because we are focused on proteins, which are the functional units of the cell, and their interactions. There have been a lot of efforts involving gene sequencing of tumors, which is valuable for identifying genes that are prevalent in cancers. Yet sequencing all the genes in tumor tissue to find a cure for cancer hasn’t worked because of cancer heterogeneity.
When you compare one patient to another, there is very little overlap with respect to genetic mutations. We decided to explore a different avenue by studying proteins and complex networks to generate the functional map of a cancer cell. Although the same genes may not be mutated, there may be commonalities in the affected pathways of cancer patients.
What is the history of your CCMI collaboration with Dr. Trey Ideker?
Trey and I have worked together for a long time. Our first interaction was working on the model organism of budding yeast. In many ways, yeast laid the framework for systems biology, specifically how to put together and visualize data.
Through our study, we realized that the most powerful type of data was networks of protein and gene interactions. When you combine these two types of information to investigate diseases and biological questions, you get a greater level of understanding.
Now, using tools like CRISPR, we can investigate interactions in mammalian cells and their role in cancer. For example, we recently described the utility of CRISPR-Cas9 and CRISPR interference screens to examine the effects of pairs of mutations in human cell lines.
How will the CCMI and your studies of biological networks ultimately help patients?
The cancer cell maps we are generating will help stratify cancer patients into groups that are predictive of prognosis and therapeutic responses. Using unbiased approaches to characterize protein interactions, the CCMI will also identify new therapeutic targets that wouldn’t be predicted based on current information in the field.