Genome Study Links Misfolded DNA to Brain Tumors
, by NCI Staff
Researchers studying brain tumors have identified a previously unknown genetic mechanism that may contribute to cancer. Their findings suggest that a change in how DNA is arranged, or packaged, in the cell nucleus may inappropriately activate a gene associated with brain cancer.
Bradley Bernstein, M.D., Ph.D., of the Broad Institute in Cambridge, MA, and his colleagues published their findings December 23 in Nature. The study focused on brain tumors known as gliomas and included lower-grade gliomas, which frequently have mutations in isocitrate dehydrogenase (IDH) genes.
IDH gene mutations in gliomas may increase the number of chemical “tags” known as methyl groups that are attached to DNA at locations throughout the genome. The new findings suggest that this process, called hypermethylation, may disrupt the packaging of DNA in the cell nucleus and lead to the activation of growth-promoting genes.
“This study identified a new mechanism that can cause cancer and helps us to better understand brain tumors,” said Dr. Bernstein, who runs a laboratory at Massachusetts General Hospital in Boston. “We now have an opportunity to see if there are analogous mechanisms in other cancers.”
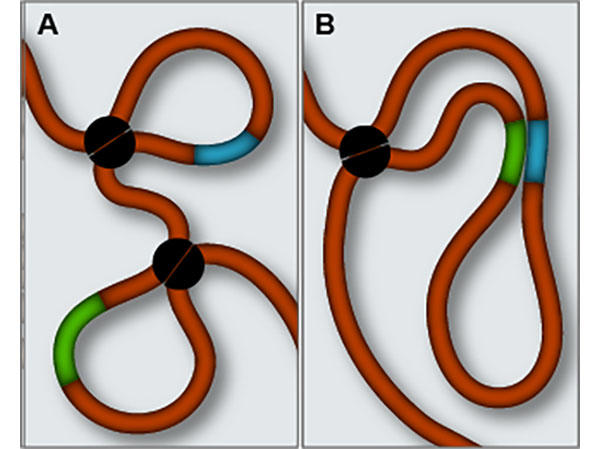
The human genome would measure about 6 feet in length if it were stretched out end to end. Cells physically arrange DNA and proteins—known as chromatin—in the space of the nucleus by forming thousands of loop-like structures. Dr. Bernstein likened these loops to that of a tied shoelace.
In gliomas with IDH mutations, however, the researchers found that some loops are misfolded. One misfolded loop, in particular, brought into close proximity two genes that usually reside far apart in the genome and rarely interact—a regulatory gene called FIP1L1 and a growth-promoting gene called PDGFRA, which has been linked to the development of gliomas.
When these genes come in close contact, a regulatory region of the FIP1L1 gene continually activates the PDGFRA gene, causing it to spur cell growth, according to the researchers. They used genomic data from projects such as The Cancer Genome Atlas (TCGA) to identify pairs of genes—including these two—that were abnormally expressed in gliomas with IDH mutations.
Another Cancer-Causing Mechanism
Cancers are frequently driven by genes that are switched on inappropriately through various mechanisms, such as mutations and fusions with DNA from other parts of the genome. But this study is the first to provide evidence that changes in the packaging of chromatin may also be a mechanism for the activation of growth-promoting genes.
Other types of cancer beyond gliomas have frequent IDH mutations, including some forms of leukemia and lymphoma. The study authors have begun to repeat their analyses in other types of cancer.
“This is an extremely interesting study,” commented Jean-Claude Zenklusen, Ph.D., of NCI’s Center for Cancer Genomics and the director of TCGA. “It expands our understanding of the complexity of processes involved in the development of cancer.”
The structure of chromatin is “exceptionally intricate,” wrote Matthew R. Grimmer, Ph.D., and Joseph F. Costello, Ph.D., of the University of California, San Francisco, in an accompanying editorial. DNA segments known as insulators normally ensure that genes and genetic elements in one loop do not interact with counterparts in other loops. But when insulators are altered or missing, which can occur through hypermethylation, the result may be inappropriate gene activity, as was observed in IDH-mutant gliomas.
Although FIP1L1 and PDGFRA may cooperate in causing cancer in IDH-mutant gliomas, neither gene has any mutations or alterations, observed coauthor William Flavahan, Ph.D., of the Broad Institute. “Our findings show that defects in chromatin structure can cause the activation of cancer-causing genes that are genetically intact,” he added.
Building on the Past
The current work builds on a large-scale genomic study published in 2008 that first linked mutations in IDH genes to brain cancer and showed that gliomas with mutations in the IDH1 gene were associated with a better prognosis compared with gliomas that lacked the mutations.
Drugs that inhibit the effects of IDH mutations have been evaluated in clinical trials, and the new findings offer some “intriguing insights” into potential therapeutic approaches, the study authors noted in a release. These include IDH inhibitors “as well as agents that reduce the associated DNA methylation or target the downstream cancer genes,” the authors wrote.
More research is needed to explore basic questions raised by the study. For instance, are there other cancer-causing genes affected by the structural changes induced by IDH1 mutations? This is likely to be the case, but many newly activated genes “may also be ‘passenger’ events, which have no functional consequences,” cautioned Drs. Grimmer and Costello in their editorial.
The new findings also illustrate the enduring value of genome sequencing data produced by TCGA, noted Dr. Zenklusen.
“TCGA data allows researchers to investigate facets of cancer biology that were not even imagined when the project began to generate information years ago,” he said. “The richness of this resource will give investigators opportunities to explore fundamental questions about cancer for decades to come.”