Novel Treatment Approach Shrinks Ovarian Tumors in Mice
, by NCI Staff
Researchers have developed a new approach for treating tumors that express mutant versions of the p53 protein, which are present in more than half of all cancers. The approach, the research team believes, could be especially beneficial in women with an aggressive and common subtype of ovarian cancer called high-grade serous.
Approximately 80 percent of ovarian cancers are the high-grade serous subtype, and nearly all tumors of this subtype have mutant forms of p53. Patients with this subtype usually aren’t diagnosed until the cancer is advanced, so long-term survival rates are low, with fewer than half of women alive 5 years after diagnosis.
In several different mouse models of high-grade serous ovarian cancer, including one that closely mimics how the cancer behaves in humans, this new therapeutic approach shrank tumors by as much as 90 percent, with no evidence of side effects.
The findings were published December 31, 2015, in Cancer Cell.
Breaking Up a Protein Gathering
The tumor suppressor protein p53 serves a critical role in preventing tumor formation. But some mutations in the TP53 gene can inactivate the p53 protein, and to date researchers haven’t been able to develop drugs that can successfully reactivate it.
An interdisciplinary team of researchers from UCLA believes they may have found a way, at least in some cases, to restore mutant p53’s tumor-suppressor functions. Their approach focuses on cases where mutant p53 clumps together into what are called amyloid aggregates.
In other diseases in which different protein aggregates commonly form, such as Alzheimer’s and Parkinson’s, this aggregation is thought to promote cell killing and directly contribute to the disease.
The team’s work builds on previous studies, which showed that some mutant versions of p53 can form protein aggregates, explained the study’s lead author, Alice Soragni, Ph.D., of the UCLA-DOE Institute.
So, adopting an approach developed by study coauthor David Eisenberg to design new therapies for diseases like Alzheimer’s, the researchers set out to see whether they could design a therapy that could stop mutant p53 from clumping together and, in doing so, restore its function.
Shifting the Equilibrium
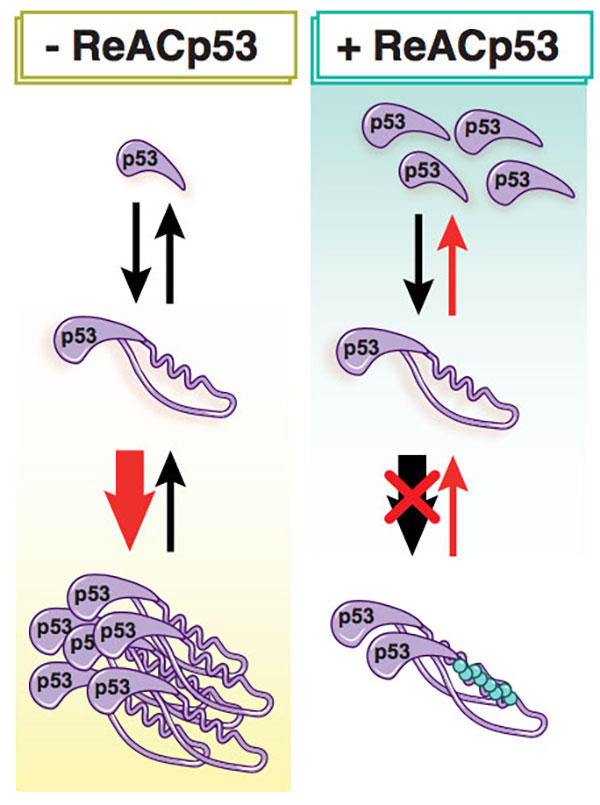
The team first showed that some mutated forms of p53 have sticky spots at specific points on their surface. These spots act like a zipper, Dr. Soragni said, so when proteins with these spots come across other p53 proteins with corresponding sticky spots, they join together.
Using the same methods they used to identify the sticky protein segments, the researchers then designed a therapy—a small portion of a protein, or peptide, that they called ReACp53—that can enter cells and block this aggregation.
In several cell-based models of mutant p53 ovarian cancer—including three dimensional models called organoids made with cells taken directly from patients—treatment with ReACp53 led to extensive cancer cell death. The peptide had no effect in ovarian cancer models with normal p53 or in models that did not express the protein.
In animal models of high-grade serous ovarian cancer, treatment with the peptide caused both primary tumors and metastatic tumors to shrink substantially. In mice that received a control treatment, tumors grew to more than twice their original size.
Even though the protein is still mutated, blocking aggregation appears to have “shifted the equilibrium” of the proteins toward a state where they can carry out their normal tumor-suppressor functions, said study coauthor Sanaz Memarzadeh, M.D., Ph.D., of the UCLA Jonsson Comprehensive Cancer Center.
“How many and what percentage of p53 mutations it will work on is something we’re still investigating, because that’s a very important question,” she said.
Further Investigation
Joanna Watson, Ph.D., of NCI’s Division of Cancer Biology, called the findings “impressive.” The responses in the various laboratory models are consistent with what would be expected when p53 is functioning normally, she said. But the data do not rule out other factors that could be responsible as well, she cautioned.
The UCLA group is not alone in its efforts. Other researchers are investigating ways to stabilize mutant p53, Dr. Watson added.
“Given that p53 is mutated in a multitude of tumors, developing a therapy that successfully restores p53’s tumor-suppressive functions to a mutant protein would be highly significant,” she said.
One of the research team’s specific goals is to define biomarkers of response to ReACp53, Dr. Memarzadeh said. “Out of the thousands of mutations in p53, we want to know which ones the drug works on, because clearly there is a subset that has these classic aggregating mutations.”
ReACp53 has been licensed to a biotech company, ADRx, Inc., co-founded by Dr. Eisenberg. The team is studying the therapy in other tumor types in which p53 is frequently mutated, Dr. Soragni said, as well as whether it may be more effective in combination with other therapies.