Genome Study Points to New Subtypes of Esophageal Cancer
, by NCI Staff
In a large-scale genomic study, researchers have characterized the genetic alterations in more than 160 cancers of the esophagus, the tube that connects the mouth and throat to the stomach.
Their findings indicate that cancers of the esophagus include multiple molecular subtypes with distinct genetic and molecular changes. Some of these subtypes may more closely resemble tumors in nearby organs than other esophageal tumors.
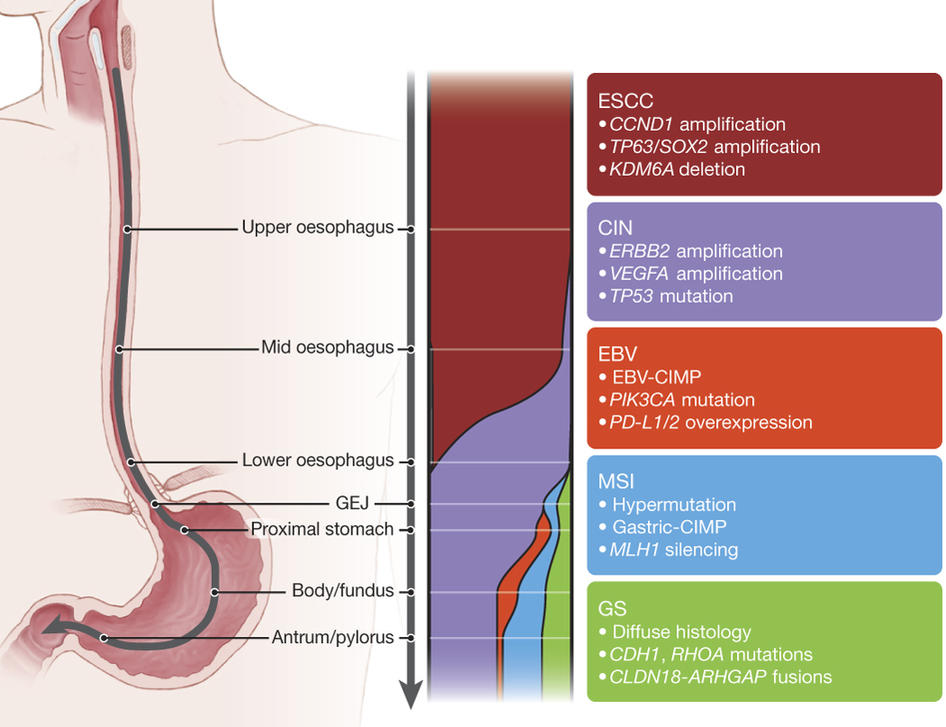
For instance, some cancers in the upper part of the esophagus were more genetically similar to cancers of the head and neck than to those in the lower part of the esophagus. And some cancers in the lower part of the esophagus were much more similar to cancers of the upper stomach than to those in the upper part of the esophagus.
The findings can help to improve the genetic and molecular classification of esophageal cancers and may also provide a framework for exploring new treatments, investigators with The Cancer Genome Atlas (TCGA) Research Network reported in Nature on January 4.
Confirming and Expanding Knowledge
“The goal of the study was to learn as much as possible about this cancer,” said Jean Claude Zenklusen, Ph.D., director of NCI’s TCGA program office. Esophageal cancers are often detected in the late stages, when they are more difficult to treat, and many do not respond to current therapies, he noted.
Esophageal cancer is rare in the United States, but it is the eighth most frequently diagnosed cancer worldwide. The most common types of esophageal cancer are adenocarcinoma and squamous cell carcinoma. Adenocarcinoma begins in cells that make and release mucus and other fluids, and squamous cell carcinoma begins in flat cells lining the esophagus.
The new findings provide evidence that esophageal squamous cell carcinoma and esophageal adenocarcinoma are distinct diseases driven by different types of genetic alterations, the study authors wrote. For instance, certain genes, such as SOX2 and TP63, were frequently present in extra copies, or amplified, in squamous cell carcinomas, whereas other genes, such as ERBB2 and VEGF, were amplified in adenocarcinomas.
“The findings confirm what some researchers have thought about esophageal cancer, but until now we have not had enough tumor samples from patients to undertake this type of integrated analysis,” said Dr. Zenklusen.
Finding Similarities
TCGA researchers began by identifying the genetic and molecular alterations in tumor samples from 164 esophageal cancers and comparing them with alterations in 359 stomach cancers and 275 head and neck cancers. The analysis included sequencing tumor DNA, analyzing chemical changes to DNA known as methylation, and checking for extra copies of chromosome regions.
The researchers identified three molecular subtypes of esophageal squamous cell carcinoma. Based on genetic mutations and patterns of gene expression, for example, the research team identified a subtype they called ESCC1 that resembles squamous cell carcinomas of the head and neck.
The researchers also observed a striking similarity between esophageal adenocarcinomas and a subtype of stomach cancer characterized by a genetic change known as chromosomal instability. These stomach cancers, which are found in the area of the stomach closest to the esophagus, were described in a 2014 TCGA study.
Based on their genomic profiles, these esophageal and stomach tumors appear to be essentially “the same cancer in organs that are next to one another,” said Dr. Zenklusen.
Implications for Treatment
TCGA researchers identified genomic changes in some esophageal tumors that could provide targets for existing targeted therapies. For instance, one-third of the esophageal adenocarcinomas had alterations in the gene ERBB2; these tumors might be good candidates for the drug trastuzumab (Herceptin®), which acts against cancers that express high levels of the protein product of this gene, HER2.
The study results also suggest that esophageal squamous cell carcinoma and esophageal adenocarcinoma “should be separated in the clinical setting so that each can be targeted according to its specific genomic features,” noted Noah D. Peyser, Ph.D., and Jennifer R. Grandis, M.D., of the University of California, San Francisco, in an accompanying editorial.
“We now understand that cancers that develop in the same location can vary greatly at the molecular level,” Drs. Peyser and Grandis wrote. “Treatment decisions made without this new wealth of genomic data can lead to patients receiving a drug that is unlikely to be beneficial, or not receiving one predicted to be effective.”
Beyond the Esophagus
The study results “add several layers of depth and sophistication to our current understanding of esophageal cancer genomics,” coauthor Adam Bass, M.D., of the Dana-Farber Cancer Institute, said in a statement. “Our hope is this work settles several long-standing uncertainties in the esophageal cancer field and will serve as the definitive reference manual for researchers and drug developers seeking more effective clinical trials and new treatment approaches.”
TCGA researchers are planning additional studies that could help further clarify relationships among esophageal cancer and other cancers.
TCGA’s final project, called the Pan-Cancer Atlas, will explore some major unanswered questions about cancer by analyzing all of the research network’s data sets. The Pan-Cancer Atlas will include a Pan-Gastrointestinal (GI) cancer analysis that will show how cancers throughout the whole GI tract relate to each other on a molecular level.
The current study suggests there may be “a continuum” of genetic alterations related to the geography of the esophagus, noted Dr. Zenklusen. “In the Pan-Cancer study, we will explore the geography of cancers of the entire gastrointestinal tract, from the esophagus to the rectum.”