Device Uses Cancer Cells' Mass to Predict Response to Treatment
, by NCI Staff
Scientists have designed a device that can detect minuscule changes in cell mass and may allow researchers to predict how cancer cells will respond to drug treatment. Such a device could potentially help clinicians determine personalized treatment regimens for individual patients, the study authors believe.
Using cancer cells from patients and mice, the researchers showed that the device, which measures changes in the mass of single cells, correctly predicted whether the cells were sensitive or resistant to a particular drug.
The research team, led by Scott Manalis, Ph.D., of the Massachusetts Institute of Technology, and David Weinstock, M.D., and Keith Ligon, M.D., of Dana-Farber Cancer Institute, also demonstrated that the device could be used to rapidly test the effects of several different drugs on cancer cells taken directly from a patient.
Although the study demonstrates the capabilities of their device, a "substantial amount of work" is still needed to determine if it can potentially guide the selection of personalized cancer treatments in the clinic, the study authors wrote.
The results of the study, which was funded in part by NCI's Innovative Molecular Analysis Technologies (IMAT) program, appeared in Nature Biotechnology on October 10.
Measuring Single-Cell Mass
Cancer treatment is shifting away from a "one-size-fits-all" approach and toward a precision medicine approach, in which therapy is selected for each patient based, in part, on the genetic and molecular makeup of the patient's cancer.
In some cases, the presence of specific genetic mutations in a tumor may help predict whether the patient is likely to respond to treatment with a particular therapy. Some researchers are trying to pinpoint these genetic mutations for diverse cancer types and to develop tests that can reliably identify them. "But for more than 90% of patients, their cancer does not have a mutation that suggests [an effective] drug," said Dr. Manalis.
An alternative approach to guiding personalized treatment, he explained, is to collect living cancer cells taken from patients and measure their biological responses to different possible therapies. Knowledge of how the patient's cancer cells respond to drug treatment in the laboratory, he continued, may better inform treatment selection.
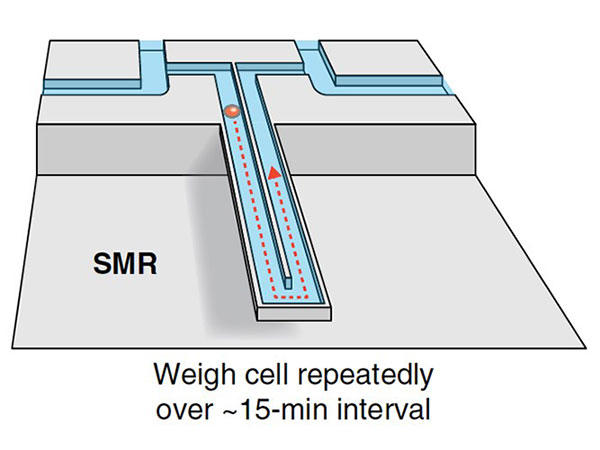
For over a decade, Dr. Manalis' team at MIT has been developing a device to help do just that. The device, called a suspended microchannel resonator (SMR), contains a series of oscillating cantilevers with hollowed-out channels through which fluids and cells are flowed. The cantilever structure, which is akin to a diving board, is “used in all kinds of technologies and as a building component,” said Tony Dickherber, Ph.D., director of the IMAT program.
Inside the SMR, Dr. Dickherber explained, the mass of a cell traveling through a channel affects how fast the cantilever wobbles. From this speed, the mass of each individual cell can be calculated in femtograms—a unit of mass one quadrillion times smaller than a gram (a paperclip has a mass of about 1 gram).
As each cell passes repeatedly through the device, its change in mass over a period of several minutes (called mass accumulation rate, or MAR) is also calculated.
"Cells are constantly taking in and secreting a lot of different things," said Dr. Manalis. "The difference between total uptake and total secretion is the mass accumulation rate," he continued, which can be indicative of cell growth.
Earlier work done by Dr. Manalis' group demonstrated that most growing cells had positive MARs and MARs of most dying cells were near zero, or in some cases, negative. For this study, the research team wanted to determine if measuring MAR following cancer treatment could rapidly and accurately predict cancer cell death or growth.
Predicting Cancer Cell Responses
First, the researchers engineered a mouse cell line to be sensitive or resistant to the targeted drug imatinib (Gleevec ®). Then they treated the cells with imatinib and measured the MARs. They found that the SMR correctly predicted the expected responses: Compared with untreated cells, the imatinib-sensitive cells had MARs near zero while the MARs of imatinib-resistant cells were unchanged.
Likewise, they saw similar results in tests of the device using different glioblastoma cell lines known to be sensitive or resistant to an experimental targeted therapy. The results were also consistent with the results of a traditional cell death test that showed whether the cells were killed by the treatment.
The device also correctly predicted responses using cells from mice engineered to have leukemia that is sensitive or resistant to imatinib, and even worked when they used only a few leukemia cells obtained from a single drop of blood. This is an important characteristic, the study authors wrote, because usually only a limited amount of cancer cells are available from patient biopsies.
Finally, the researchers tested the device using clinical samples from human patients. They found that they could quickly measure MARs of acute myeloid leukemia (AML) cells that were obtained directly from a patient and then treated with several drugs in the lab. They also used the SMR to detect changes in the MAR of leukemia cells before and after a patient received treatment.
Advantages and Challenges
Throughout their experiments, the researchers found that MARs of individual cells from the same tumor varied widely, which, they wrote, reflects tumor heterogeneity—diversity in the molecular makeup and behavior of cancer cells in a tumor.
Other tests of drug sensitivity commonly used in the lab measure the average growth or death rate of a group of cells, but that average may not represent the true behavior of the entire cell population, Dr. Manalis said. Tumor heterogeneity has been linked to drug resistance, so being able to measure that heterogeneity in lab tests could potentially produce more accurate predictions of tumor responses.
The device may be able to overcome other shortcomings of existing drug sensitivity tests in that the SMR can quickly and accurately give readings using a small number of cells, said Dr. Dickherber. This short timeline is a major advantage, said Dr. Manalis, especially when testing cells that have been taken from a patient because "the sooner you can measure them, the closer the cells are likely to be to their original state."
But there are limitations to the new device as well, Dr. Dickherber cautioned.
For instance, the cell culture procedure required to treat patients' cells before they are analyzed by the SMR may introduce unnatural biological changes that can skew results. And the researchers don’t yet know whether they will be able to use the device to test cells from patients with solid tumors because of potential damage to the cells while they are being prepared for the SMR.
Dr. Manalis believes that another "useful capability" of their device is that cells tested with the SMR can still be used for additional analyses, including sequencing. For example, Dr. Manalis and his team are performing single-cell gene expression measurements after they acquire MAR measurements in order to better understand what makes some cells responsive and others resistant to the same drug treatment.